Mean Imputation for Missing Data (Example in R & SPSS)
Let’s be very clear on this: Mean imputation is awful!
Do you think about using mean imputation yourself? Stop it NOW!
Sorry for the drama, but you will find out soon, why I’m so much against mean imputation. First, let me define what we are talking about.
Mean imputation (or mean substitution) replaces missing values of a certain variable by the mean of non-missing cases of that variable.
Sounds easy to apply, doesn’t it? So why is it so evil to use mean substitution? Click on the buttons below to select the topic you are interested in:
Mean Imputation Pros & Cons Mean Imputation in R (Example) Mean Imputation in SPSS (Video) Ask me a Question (It's Free)
Advantages and Drawbacks of Mean Substitution
You probably already noticed that I’m not a big fan of mean imputation. However, I’ll be fair and show you also the advantages of the method:
- Missing values in your data do not reduce your sample size, as it would be the case with listwise deletion (the default of many statistical software packages, e.g. R, Stata, SAS or SPSS). Since mean imputation replaces all missing values, you can keep your whole database.
- Mean imputation is very simple to understand and to apply (more on that later in the R and SPSS examples). You can explain the imputation method easily to your audience and everybody with basic knowledge in statistics will get what you’ve done.
- If the response mechanism is MCAR, the sample mean of your variable is not biased. Mean substitution might be a valid approach, in case that the univariate average of your variables is the only metric your are interested in.
We learned some reasons why mean imputation is so popular among data users. However, let’s move on to the more important part – the drawbacks of mean imputation:
- Mean substitution leads to bias in multivariate estimates such as correlation or regression coefficients. Values that are imputed by a variable’s mean have, in general, a correlation of zero with other variables. Relationships between variables are therefore biased toward zero.
- Standard errors and variance of imputed variables are biased. For instance, let’s assume that we would like to calculate the standard error of a mean estimation of an imputed variable. Since all imputed values are exactly the mean of our variable, we would be too sure about the correctness of our mean estimate. In other words, the confidence interval around the point estimation of our mean would be too narrow.
- If the response mechanism is MAR or MNAR, even the sample mean of your variable is biased (compare that with point 3 above). Assume that you want to estimate the mean of a population’s income and people with high income are less likely to respond; Your estimate of the mean income would be biased downwards.
In summary: There are a few advantages, but many serious drawbacks. On top of that, we can also benefit from the advantages with more advanced imputation methods (e.g. predictive mean matching or stochastic regression imputation). To make it short, there is basically no excuse for using mean imputation.
In the following step-by-step example in R, I’ll show you how mean imputation affects your data in practice.
Mean Imputation in R (Example)
Before we can start with the example, we need some data with missing values. Let’s create some ourself:
##### Create some synthetic data with missings ##### set.seed(87654) # Reproducibility N <- 1000 # Sample size # Some random variables x1 <- round(rnorm(N), 2) x2 <- round(x1 + rnorm(N, 10, 5)) x3 <- round(runif(N, -100, 20)) # Insert missing values x1[rbinom(N, 1, 0.2) == 1] <- NA # 20% missingness x2[rbinom(N, 1, 0.05) == 1] <- NA # 5% missingness x3[rbinom(N, 1, 0.7) == 1] <- NA # 70% missingness # Indicator for missings (needed later) x1_miss_ind <- is.na(x1) x2_miss_ind <- is.na(x2) x3_miss_ind <- is.na(x3) # Store variables in a data frame data <- data.frame(x1, x2, x3) head(data) # First 6 rows of our data
Our data consists of the three variables X1, X2, and X3 – all of them have missing values (i.e. NAs). This is how the first 6 rows of our example data look like:
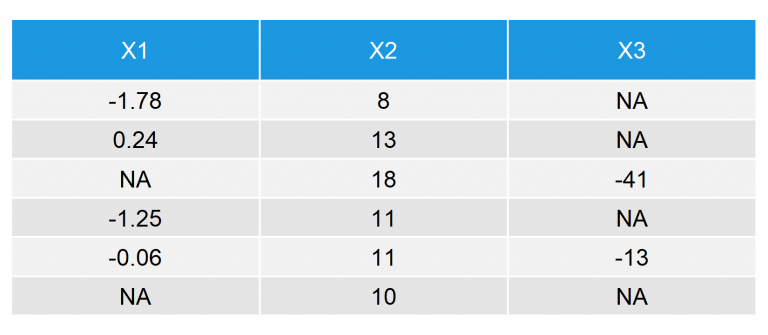
Table 1: First 6 Rows of Our Example Data for Mean Imputation
Mean Imputation of One Column
Let’s move on to the part we are interested in: The mean imputation. If we want to impute only one column of our data frame, we can use the following R code:
##### Imputation of one column (i.e. a vector) ##### data$x1[is.na(data$x1)] <- mean(data$x1, na.rm = TRUE)
That’s it – plain and simple. So, what is this code doing exactly?
- data$x1 tells R to use only the column x1.
- is.na() is a function that identifies missing values in x1. (More infos…)
- The squared brackets [] tell R to use only the values where is.na() == TRUE, i.e. where x1 is missing.
- <- is the typical assignment operator that is used in R.
- mean() is a function that calculates the mean of x1.
- na.rm = TRUE specifies within the function mean() that missing values should not be used for the mean calculation (na.rm = FALSE would be impossible and would lead to an error).
Mean Imputation of Multiple Columns
Often we want to impute all data at once. In R, that is easily possible with a for loop.
##### Imputation of multiple columns (i.e. the whole data frame) ##### for(i in 1:ncol(data)) { data[ , i][is.na(data[ , i])] <- mean(data[ , i], na.rm = TRUE) } head(data) # Check first 6 rows after substitution by mean
With our for loop, we iterate along all columns of our data and apply to each column the same operation as in the previous example, in which we imputed only one column. By doing so, we can impute the whole database with 3 lines of code.
Evaluation of Imputed Values
As I told you, mean imputation screws your data. I’ll show you graphically what I’m talking about:
##### Density of x1 pre and post imputation ##### # Density of observed data plot(density(data$x1[x1_miss_ind == FALSE]), xlim = c(- 4, 4), ylim = c(0, 0.9), lwd = 2, main = "Density Pre and Post Mean Imputation", xlab = "X1") # Density of observed & imputed data points(density(data$x1), lwd = 2, type = "l", col = "red") # Legend legend("topleft", c("Before Imputation", "After Imputation"), lty = 1, lwd = 2, col = c("black", "red"))
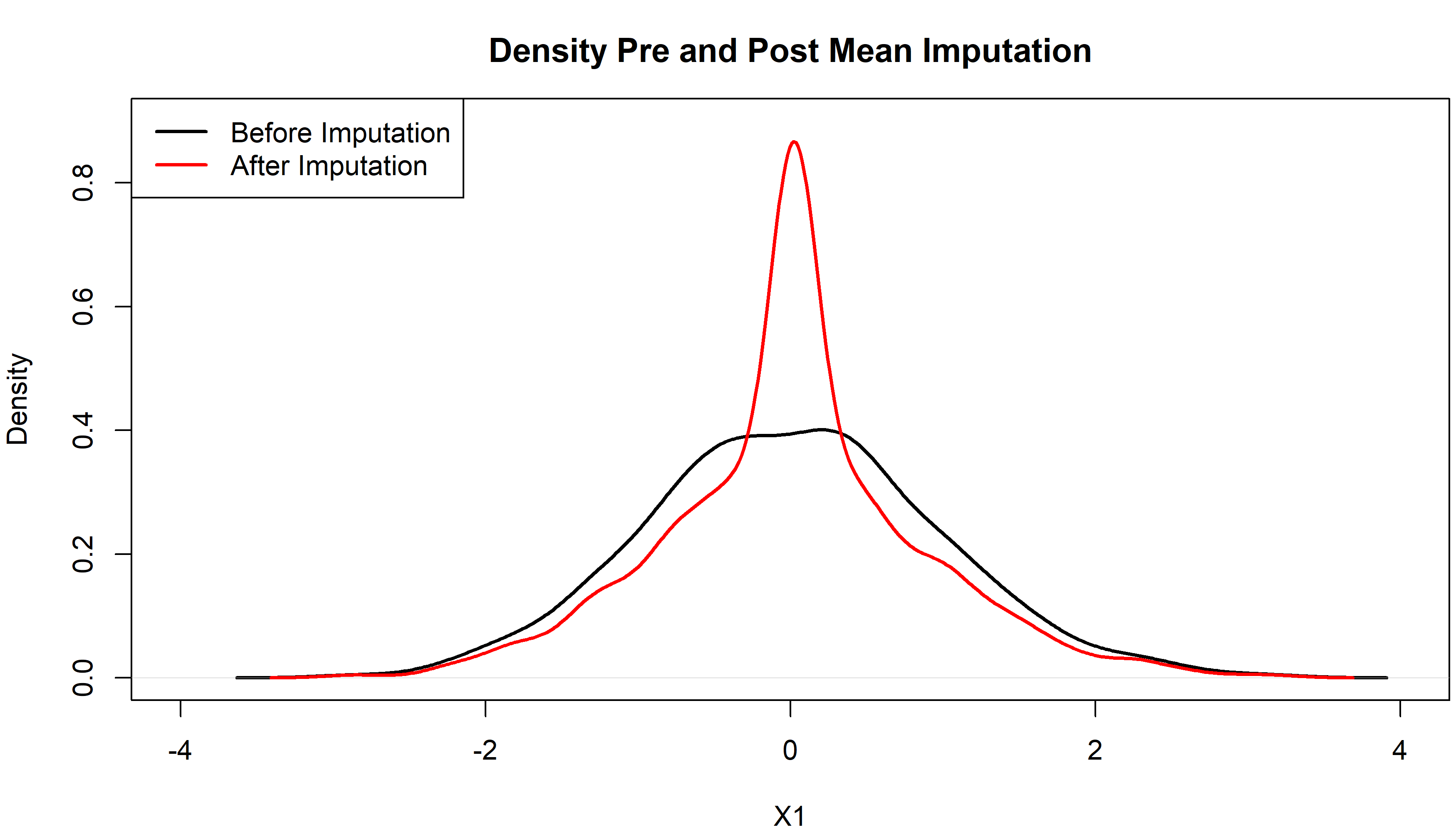
Figure 1: Density of X1 Pre and Post Mean Imputation
Figure 1 displays the density of X1 before (in black) and after (in red) the imputation. Before imputation, X1 is following a normal distribution. After imputing the mean, however, our density has a weird peak at zero (in our example the mean of X1).
So, how does that affect our data analysis? Let’s do some univariate descriptive statistics:
##### Descriptive statistics for X1 ##### # Pre imputation round(summary(data$x1[x1_miss_ind == FALSE]), 2) ### Min. 1st Qu. Median Mean 3rd Qu. Max. ### -2.95 -0.64 0.00 0.02 0.64 3.23 # Post imputation round(summary(data$x1), 2) ### Min. 1st Qu. Median Mean 3rd Qu. Max. ### -2.95 -0.45 0.02 0.02 0.45 3.23
The mean before and after imputation is exactly the same – no surprise. Since our missing data is MCAR, our mean estimation is not biased.
The problem is revealed by comparing the 1st and 3rd quartile of X1 pre and post imputation.
First quartile before and after imputation: -0.64 vs. -0.45.
Third quartile before and after imputation: 0.64 vs. 0.45.
Both quartiles are shifted toward zero, after substituting missing data by the mean. In other words, the quartiles are highly biased.
Even bigger problems arise for multivariate measures. For instance, let’s evaluate the correlation of X1 and X2:
##### Correlation of X1 and X2 ##### # Pre imputation round(cor(data$x1[x1_miss_ind == FALSE & x2_miss_ind == FALSE], data$x2[x1_miss_ind == FALSE & x2_miss_ind == FALSE]), 3) ### 0.268 # Post imputation round(cor(data$x1, data$x2), 3) ### 0.238
Again, we observe bias after imputation. The correlation coefficient between X1 and X2 is shifted toward zero.
We can also observe that graphically:
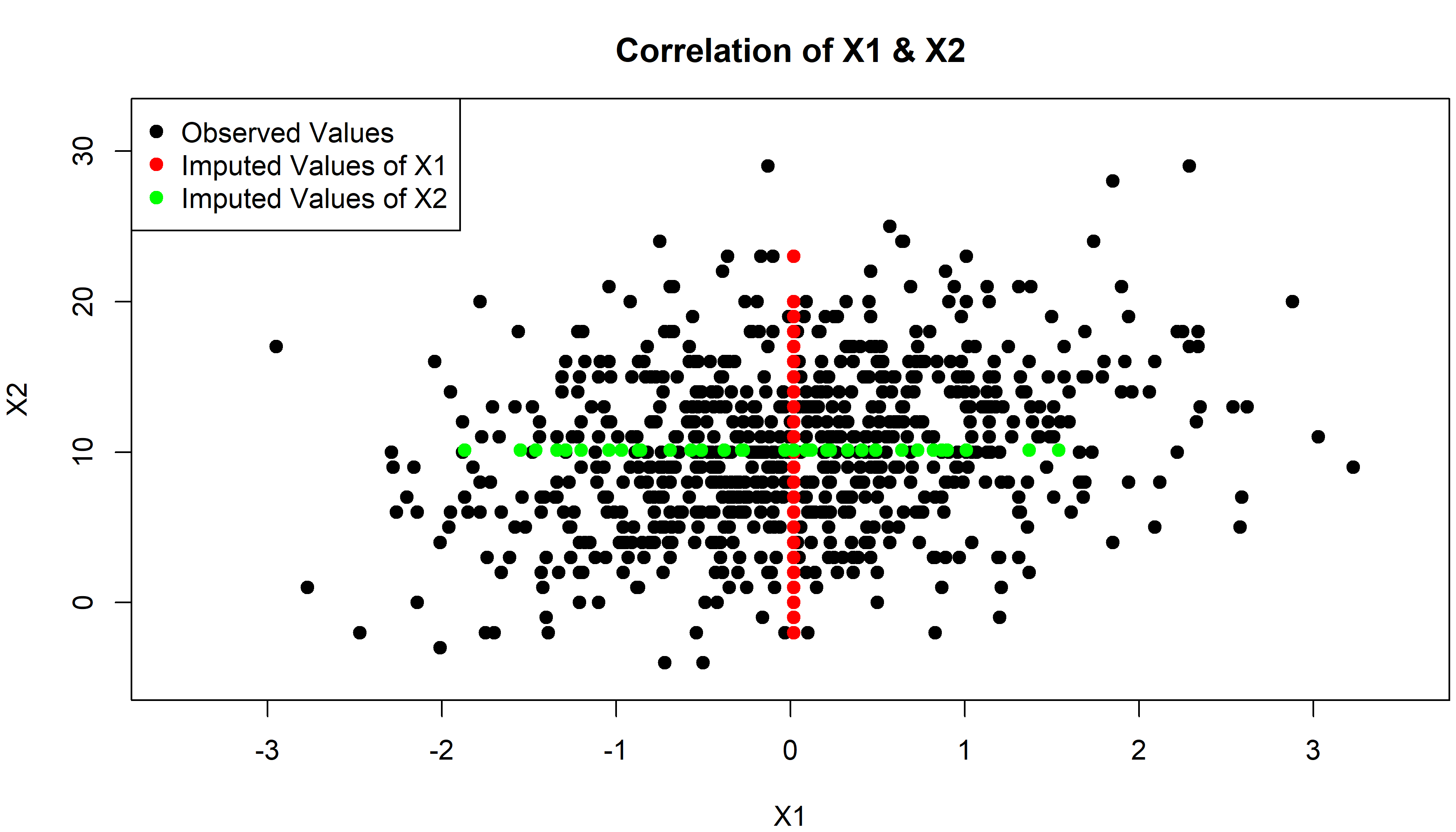
Figure 2: Correlation Plot of X1 & X2 After Mean Imputation
Figure 2 illustrates the correlation between X1 and X2 for observed and imputed data. Observed values are shown in black, imputed values of X1 in red, and imputed values of X2 in green.
The observed values are widely spread with a small positive correlation. However, this distribution of X1 and X2 is not reflected by the imputed values. Since all missing values of X1 and X2 were imputed by each variable’s average, imputed and observed values are not correlated.
Imputation of Row Means
A less known modification of mean imputation – about which we haven’t talked yet – is an imputation by row means. Instead of imputing the mean of a column (as we did before), this method computes the average of each row.
Imputing the row mean is mainly used in sociological or psychological research, where data sets often consist of Likert scale items. In research literature, the method is therefore sometimes called person mean or average of the available items.
Row mean imputation faces similar statistical problems as the imputation by column means. However, it is also very easy to apply in R:
##### Imputation of one row (i.e. a row vector) ##### data[1, ][is.na(data[1, ])] <- mean(as.numeric(data[1, ]), na.rm = TRUE) ##### Imputation of multiple rows (i.e. the whole data frame) ##### for(i in 1:nrow(data)) { data[i, ][is.na(data[i, ])] <- mean(as.numeric(data[i, ]), na.rm = TRUE) } head(data) # Check first 6 rows after substitution by mean
Hint: If all cells of a row are missing, the method is not able to impute a value. R imputes NaN (Not a Number) for these cases.
Mean Imputation in SPSS (Video)
As one of the most often used methods for handling missing data, mean substitution is available in all common statistical software packages. If you want to learn how to conduct mean imputation in SPSS, I can recommend the following YouTube video.
Based on some example data, the speaker Todd Grande explains how to apply mean imputation in SPSS. He also speaks about the impact of listwise deletion on your data analysis and compares this deletion method with mean imputation (see also the first advantage of mean imputation I described above).
Now It’s On You!
I showed you in this article why mean imputation screws the quality of your data analysis.
Now I’d like to hear from you!
Have you already used mean substitution in the past? Would you do it again nowadays? Do your colleagues or your boss share your opinion?
Write me about your experiences in the comments (of cause questions are also welcome)!
Appendix
The header graphic of this page illustrates an extreme mean substitution.
The black triangles reflect observed values – none of them close to zero. The red dots reflect imputed values – all of them exactly at zero.
Here’s the code for the graphic:
set.seed(2332332) # Seed for reproducibility par(bg = "#1b98e0") # Set background colors N <- 10000 # Sample size x <- rnorm(N) # Some random data y <- rnorm(N) x <- x[y > 0.3 | y < - 0.3] # Delete values in middle of plot y <- y[y > 0.3 | y < - 0.3] plot(x, y, pch = 17, col = "#353436") N_imp <- 500 # Add some red points at zero x_imp <- rnorm(N_imp ) y_imp <- rep(0, N_imp) points(x_imp , y_imp, pch = 20, col = "brown3")
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.







10 Comments. Leave new
Can you share the pros and cons of Hot deck imputation?. Also, can you brief on mean versus conditional mean and Hot deck imputation versus Conditional Hot deck
Hi indu,
The main pro of Hot Deck imputation is that it imputes values that where observed for other individuals. Therefore, the imputed values are supposed to be more realistic.
If you want to learn more about Hot Deck imputation, I can recommend to have a look at this paper of Andridge & Little.
I hope that helps.
Joachim
I haven’t found any instructions/syntax on how to replace a missing value with the value of another variable for the same case in SPSS.
I need to replace missing values with the same person’s average on the other items that make up a sum variable. The variables are Likert scored items. I have very little missing values (about .3%), but I need to have none, as I have to use AMOS later.
Please can you help me?
Hi Sussa,
Thank you for your comment.
Unfortunately, I am not an expert for SPSS syntax. In R, you could do something like that:
Whereby x1 would be your item with missing values and x2, x3, and x4 would be the observed items.
You can use AGGREGATE to add the mean or other statistics for variables to be imputed to the dataset and then use a simple IF or DO IF to replace the missing value
Hey Jon,
Thanks a lot for your help, that’s very kind of you!
Regards,
Joachim
Hi!
I was wondering what the standard percentage of missing data is in order to include the case vs deleting?
EX- if a case is missing 50% of data should it be excluded or would it be 20%?
Hey Lane,
This strongly depends on your specific variable and your field of research. This article gives some good insights.
Regards,
Joachim
Hi!
I am currently in search to see if there’s a way we can find the mean of missing values while grouping with another variable.
example: Calculate missing values of age by computing the feature mean grouped by country
Hello Vanessa,
Could you explain your example a bit more in detail? I am not sure if I got it.
Regards,
Cansu