R Replace NA with 0 (10 Examples for Data Frame, Vector & Column)
A common way to treat missing values in R is to replace NA with 0.
You will find a summary of the most popular approaches in the following.
Choose one of these approaches according to your specific needs.
What are you interested in?
Data Frame: Replace NA with 0 Vector or Column: Replace NA with 0 Is the Replacement of NA’s with 0 Legit? Alternative Ways to Handle Missing Data
R Replace NA with 0 in a Data Frame
Consider the following example data frame in R.
data <- data.frame(x1 = c(3, 7, 2, 5, 5), x2 = c(NA, 8, 6, NA, 5), x3 = c(3, NA, 5, 1, 9)) data
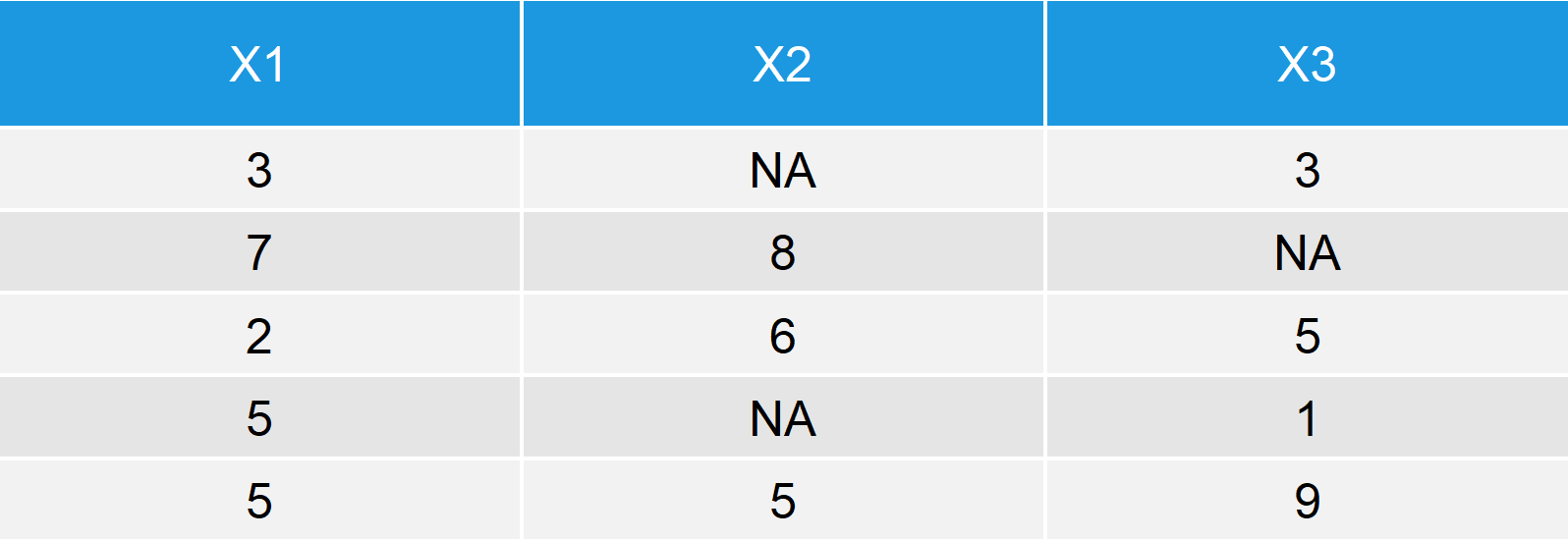
Table 1: Exemplifying Data Frame with Missing Values
I’m creating some duplicates of the data for the following examples.
data_1 <- data data_2 <- data data_3 <- data data_4 <- data data_5 <- data # Example for data frame with factor variable data_5$x3 <- as.factor(data_5$x3)
Data Frame Example 1: The Most Common Way to Replace NA with 0
data_1[is.na(data_1)] <- 0
In this video, I’m applying our is.na() approach of Example 1 to a real data set (and a vector as shown later).
Data Frame Example 2: Replace NA During the Data Export
setwd("Insert your path here") write.csv(data_2, "data_2.csv", na = "0")
Data Frame Example 3: dplyr Package
library("dplyr") data_3 <- data_3 %>% mutate(x2 = coalesce(x2, 0), x3 = coalesce(x3, 0))
Data Frame Example 4: imputeTS Package
library("imputeTS") data_4 <- na.replace(data_4, 0)
Data Frame Example 5: Database with Factor Variables
One common issue for replacing NA with 0 in an R database is the class of the variables in your data.
The previous examples work fine, as long as we are dealing with numeric or character variables.
However, if you have factor variables with missing values in your dataset, you have to do an additional step.
i <- sapply(data_5, is.factor) # Identify all factor variables in your data data_5[i] <- lapply(data_5[i], as.character) # Convert factors to character variables data_5[is.na(data_5)] <- 0 # Replace NA with 0, as shown in Example 1 data_5[i] <- lapply(data_5[i], as.factor) # Convert character columns back to factors
Insert Zeros for NA Values in an R Vector (or Column)
As you have seen in the previous examples, R replaces NA with 0 in multiple columns with only one line of code. However, we need to replace only a vector or a single column of our database. Let’s find out how this works.
First, create some example vector with missing values.
vec <- c(1, 9, NA, 5, 3, NA, 8, 9) vec # Duplicate vector for later examples vec_1 <- vec vec_2 <- vec vec_3 <- vec vec_4 <- vec vec_5 <- as.factor(vec) # Example for factor vector
Vector Example 1: The Most Common Way to Replace NA in a Vector
vec_1[is.na(vec_1)] <- 0
Vector Example 2: Create Your Own Function to Replace NA’s
fun_zero <- function(vector_with_nas) {
vector_with_nas[is.na(vector_with_nas)] <- 0
return(vector_with_nas)
}
vec_2 <- fun_zero(vec_2)
fun_zero <- function(vector_with_nas) { vector_with_nas[is.na(vector_with_nas)] <- 0 return(vector_with_nas) } vec_2 <- fun_zero(vec_2)
Vector Example 3: Using the replace() Function
vec_3 <- replace(vec_3, is.na(vec_3), 0)
Vector Example 4: Using the ifelse() Function
vec_4 <- ifelse(is.na(vec_4), 0, vec_4)
Vector Example 5: Exchange NA’s with Zero in Factor Vectors
vec_5 <- as.numeric(as.character(vec_5)) # Note: Transform vec_5 as.character first, # otherwise you might lose the levels of your vector vec_5[is.na(vec_5)] <- 0 # Similar to Example 1 vec_5 <- as.factor(vec_5)
As you can see, there are many different ways in R to replace NA with 0 – All of them with their own pros and cons.
If you want to investigate even more possibilities for a zero replacement, I can recommend the following thread on stackoverflow.
Is the Replacement of NA’s with 0 Legitimate?
Beside the question how to find and replace NA with 0 in R, the question arises whether such a replacement screws our statistical data analyses.
As most of the time in statistics, the answer is: It depends! If it is meaningful to substitute NA with 0, then go ahead.
For instance, let’s say we have the item “How much did you spend for holidays last year?” and people without any spending for holidays are represented by NA. Then it would be logical to change NA to 0, since these people basically spend zero money for holidays.
However, if we have NA values due to item nonresponse, we should never replace these missing values by a fixed number, i.e. 0.
Consider the following example:
set.seed(765) # Set seed to make the example reproducible example_vector <- rnorm(10000) # Example vector: Normal distribution with 10000 observations example_vector[1:1000] <- NA # Insert missing values for the first 1000 observations plot(density(example_vector, na.rm = TRUE), ylim = c(0, 0.7), xlab = "Example Vector", main = "With & without replacement of NA with 0") example_vector[is.na(example_vector)] <- 0 # As in Example 1 in R: Replace NA with 0 lines(density(example_vector, na.rm = TRUE), col = "red") # Plot density of the example vector # after replacing NA's with 0
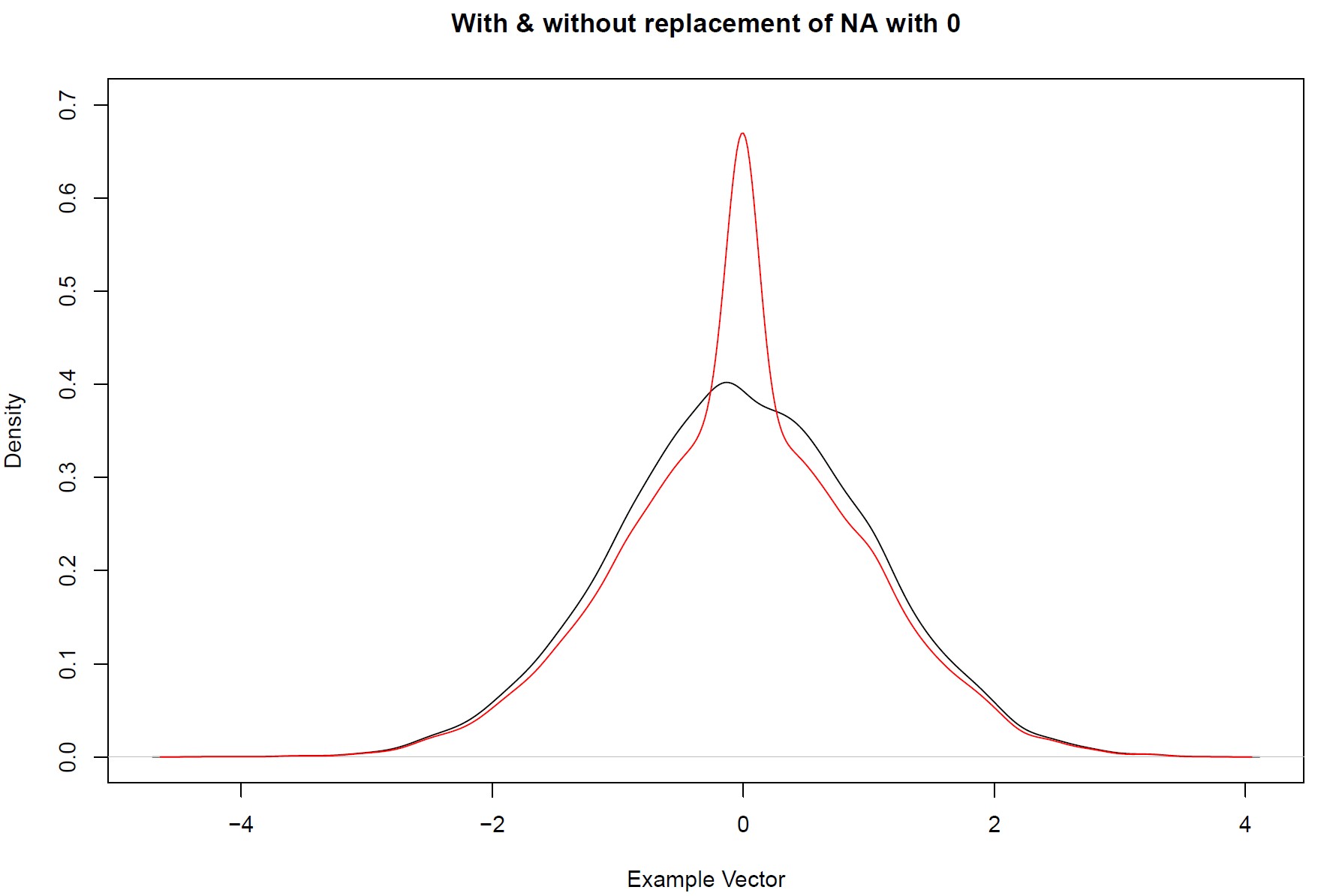
Graphic 1: R Replace NA with 0 – Densities with & without Zero-Replacement
As you can see in the example, the density of a normal distribution would be highly screwed toward zero, if we just substitute all missing values with zero (as indicated by the red density).
Alternatives to the Replacement of Missing Data by 0
The statistical analysis with missing data is a whole domain of statistical research.
The imputation of missing values is one of the most popular approaches nowadays.
When data is imputed, new values are estimated on the basis of imputation models in order to replace missing values by these estimates.
In fact, the replacement of NA’s with zero could also be considered as a very basic data imputation (zero imputation).
Another popular approach is casewise deletion (also called listwise deletion).
In casewise or listwise deletion, all observations with missing values are deleted – an easy task in R.
This approach has its own disadvantages, but it is easy to conduct and the default method in many programming languages such as R.
Conclusion
To change NA to 0 in R can be a good approach in order to get rid of missing values in your data.
The statistical software R (or RStudio) provides many ways for the replacement of NA’s.
However, such a replacement should only be conducted, if there is a logical reasoning for converting NA’s to zero.
Now it’s Your Turn!
I put together 10 different ways how to replace NA’s with 0 in R.
Now I’m interested to hear from you.
Are you handling NA’s with the popular approaches of Data Frame Example 1 and Vector Example 1? Or are you using other ways? Do you still have any issues with your NA’s?
Let me know in the comments!
References
Moritz, S. (2017). Package imputeTS
Wickham, H., Francois, R., Henry, L., Müller, K., and RStudio (2017). Package dplyr
Appendix
How to create the header graph:
The header graphic of this page shows a correlation plot of two continuous (i.e. numeric) variables, created with the package ggplot2.
The dark blue dots indicate observed values. The light blue dots indicate NA’s that were replaced by zero.
library("ggplot2") # Load R package ggplot2 set.seed(9876543) # Set seed to ensure reproducibility x1 <- rnorm(2000) # Random normally distributed x1 x2 <- 2 * x1 + rnorm(2000) # Generate x2 correlated with x1 x2[1801:2000] <- 0 # Set some values of x2 to zero data_ggp <- data.frame(x1, x2) # Store x1 and x2 in a data frame colours <- c(rep(1, 1800), rep(2, 200)) # Set colours ggp <- ggplot(data_ggp, aes(x = x1, y = x2)) + # Create ggplot geom_point(aes(col = colours , size = 1.1)) + theme(legend.position = "none") ggp
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.







12 Comments. Leave new
I simply desired to say thanks once more. I’m not certain the things that I could possibly have used without the entire aspects revealed by you over such subject matter. It seemed to be the alarming dilemma in my opinion, but discovering the very professional fashion you solved it took me to cry for fulfillment. Extremely grateful for this service as well as pray you are aware of a great job that you’re undertaking educating the others through your webblog. Most likely you have never come across all of us.
Thanks for the kind words Ahmad. I’m glad to hear that I could help you!
Thank you so so much!!! It helped
Thanks for your feedback DK, I’m glad to hear that.
This is brilliant! Thank you for taking the time to put together such a well versed set of examples.
Thank you very much Mauricio!
hello, could you provide any assistance with merging data frames. I had a look at your page about it but this particular scenario doesn’t come up.
I have 2 dataframes:
1. x y
a A1 blue
b A2 N/A
c A3 yellow
2. x y
a A1 N/A
b A2 red
c A3 N/A
each dataframe fills in the holes in the other (the N/As). I’ve tried rbind, cbind, etc… but they only seem to try and add extra columns or rows. I just want to fill in those blanks by merging the dataframes. I hope there is an easy way to do this… there is in excel, but that’s too slow.
Any guidance appreciated, but no problem if not!
Cheers
Hey Freya,
Is the following R code what you are looking for?
Please note that this only works in case both data frames contain exactly the same values in x, and in case both data frames are ordered the same way.
I hope that helps!
Joachim
Dear Joachim,
Thanks for the explanations. However, in my case, I would like to replace randomly 1000 NA values in a column with 0s. None of your examples provide an exact solution for this task. Could you please suggest one? I think it would be helpful for me and for other people as well.
Thanks
Hey Filipe,
Thank you for the kind comment.
Below, I have created an example that explains how to do that:
I hope that helps!
Regards,
Joachim
Thank you, excellent explanation
Thanks a lot Marcos, glad you think so!