Mean by Group in R (2 Examples) | dplyr Package vs. Base R
In this tutorial you’ll learn how to compute the mean by group in the R programming language.
I’ll show two different alternatives including reproducible R codes.
Let’s dig into it!
Example Data
For the following examples, I’m going to use the Iris Flower data set. Let’s load the data to R:
data(iris) # Load Iris data head(iris) # First rows of Iris
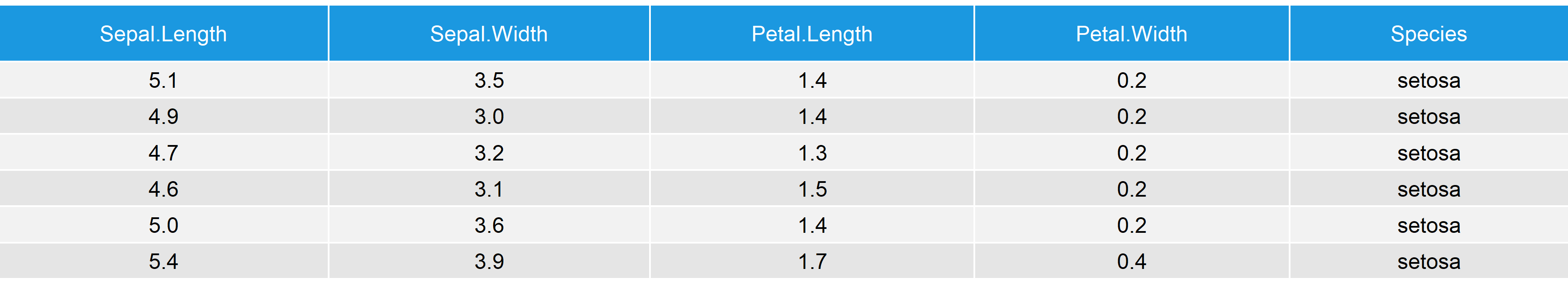
Table 1: The Iris Data Matrix.
As you can see based on Table 1, the Iris Flower data contains four numeric columns as well as the grouping factor column Species
Next, I’ll show you how to calculate the average for each of these groups. Keep on reading!
Example 1: Compute Mean by Group in R with aggregate Function
The first example shows how to calculate the mean per group with the aggregate function.
We can compute the mean for each species factor level of the Iris Flower data by applying the aggregate function as follows:
aggregate(x = iris$Sepal.Length, # Specify data column by = list(iris$Species), # Specify group indicator FUN = mean) # Specify function (i.e. mean) # Group.1 x # setosa 5.006 # versicolor 5.936 # virginica 6.588
The RStudio console output shows the mean by group: The setosa group has a mean of 5.006, the versicolor group has a mean of 5.936, and the virginica group has a mean of 6.588.
Note: By replacing the FUN argument of the aggregate function, we can also compute other metrics such as the median, the mode, the variance, or the standard deviation.
Example 2: Compute Mean by Group with dplyr Package
It’s definitely a matter of taste, but many people prefer to use the dplyr package to compute descriptive statistics such as the mean. This example shows how to get the mean by group based on the dplyr environment.
Let’s install and load the dplyr package to R:
install.packages("dplyr") # Install dplyr package library("dplyr") # Load dplyr package
Now, we can use all the functions of the dplyr package – in our case group_by and summarise_at:
iris %>% # Specify data frame group_by(Species) %>% # Specify group indicator summarise_at(vars(Sepal.Length), # Specify column list(name = mean)) # Specify function # A tibble: 3 x 2 # Species Sepal.Length # # setosa 5.01 # versicolor 5.94 # virginica 6.59
The output of the previous R syntax is a tibble instead of a data.frame. However, the results are the same as in Example 1.
Video, Further Resources & Summary
On the Statistics Globe YouTube channel, you can also find a tutorial video, where I explain the content of this topic in some more detail:
This tutorial illustrated how to compute the means for certain data frame subsets (i.e. groups) in the R programming language. In case you want to learn more about the theoretical research concept of the mean, I can recommend the following video of the mathantics YouTube channel:
Furthermore, you could also have a look at some of the related R tutorials that I have published on my website:
- aggregate Function in R
- Mean in R
- Mean Across Columns & Rows
- Weighted Mean in R
- Geometric Mean in R
- Harmonic Mean in R
- Mean of Data Frame Column
- Mean Imputation
- Median in R
- Mode in R
- R Functions List (+ Examples)
- The R Programming Language
I hope you found the tutorial helpful. However, if you have any questions or comments, don’t hesitate to let me know below.







26 Comments. Leave new
After I used the dplyr option I’ve got this warning message
funs() is soft deprecated as of dplyr 0.8.0
please use list() instead
# Before:
funs(name = f(.))
# After:
list(name = ~ f(.))
So, maybe you could update the example code as:
iris%>%
group_by(Species)%>%
summarise_at(vars(Sepal.Length), list(name=mean))
Regards!
Hey Ruben,
Thank you for the hint, I’ve just changed the code accordingly 🙂
Regards,
Joachim
Hi! I’m fairly new to using R, after finding the mean values of two categorical groups I have how would I plot those means in a bar chart? Thank you
Hi Ben,
Good question! You may have a look at this tutorial: https://statisticsglobe.com/barplot-in-r
You may use the means of your two groups as height of the bar charts, i.e. store the two means in the vector “values” as shown in Example 1.
Regards,
Joachim
Thank you so much for the easy to follow instructions! I’ve been working on project since last week for my job and this cleared everything up for me in 5 minutes! Thank you!!!
Thank you for the comment, Julianna. I’m glad to hear that it helped! 🙂
Thanks so much for this! It was very helpful 🙂
Thanks Alicia, I’m happy to hear that 🙂
Thank you for these instructions – very helpful! I am wondering if the output for the dplyr method is rounded? If so, is there a way for the output to not be rounded like the aggregate function?
Thanks!
Hi Jan,
Thank you for the comment and the kind words!
Actually, this is a very good question! The dplyr package returns the data in tibble format (in contrast to the data.frame format returned by the aggregate function). Tibbles automatically display the data rounded to two digits. However, the actual values stored in the tibble are NOT rounded.
You can see that by converting the tibble back to data.frame format:
I also found this thread, which is discussing this topic in some more detail.
I hope that helps!
Joachim
Please help,
# Calculate the average_pop and median_pop columns
counties_selected %>%
group_by(region, state) %>%
summarize(total_pop = sum(population)) %>% ungroup()%>%
summarize(average_pop = mean(population), median_pop = median(population)
)
I got “object population not found”
Hey Ngoc,
Thanks for the comment!
I need a few more details. Is counties_selected a data frame? Could you tell me the column names of counties_selected?
Regards,
Joachim
Hi Joachim, thanks for this tutorial! My question: How do I switch rows and columns? So that I for example have only have 2 columns (sex) and x rows. Thank you!
My example: I want do group all my latent variables by sex:
data %>%
group_by(sex) %>%
summarise_at(vars(example1, example2, example3, examplex), list(mean))
If I do so, I get very long rows.
and one hint: I only use “list(mean))” and it name every column correct.
Thank you in advance!
Hey,
Thank you for the nice comment!
Could you illustrate how your input data looks like? What is contained in the variables example1, example2, and so on?
Regards
Joachim
Hi Joachim,
the variables are for example: motivation, work engagement, authentic leadership
they were all measured with 5 or 7-point likert scales and I just want to list the means.
is there any syntax for swichting the rows and columns?
One more question: Where has to be placed the syntax to round the means?
i would like the means to be rounded to two decimal places.
data %>%
group_by(sex) %>%
summarise_at(vars(motivation, work engagement, authentic leadership),
list(mean), round(2))
this doesn’t work…
Hi again,
I have created some example data to illustrate how you might do that.
Example data:
Calculate mean values by group:
Convert data from wide to long format:
I hope that helps!
Joachim
Hello
am trying to find the means by group but it brings an error each time
mydata%>%
group_by(mydata$Subspecies)%>%
summarise_at(vars(mydata$Gall.sizes),list(name=mean))
str(mydata)
Hey Cynthia,
What does the error message say?
Regards
Joachim
Hello,
my columns are called frame_number, name, x,y. Is there a possibility to group x by “name” but also the “frame number” at the same time? So that I would get the mean of all x values grouped by name but also just for a specific frame number?
Thanks in advance!
Hey Remya,
You may group your data frame by multiple columns as explained here.
Regards,
Joachim
Hallo Joachim,
ich erkläre es mal auf Deutsch, weil ich mir da leichter tue. Ich möchte den Mittelwert von den “x”-Werten pro Kategorie der Spalte “name” (dort habe ich blau, rot, gelb, usw.) und pro “frame_number”. Ich habe also für jeden Frame und jede Farbe mehrere x-Werte und ich möchte den Mittelwert bilden, damit ich im Endeffekt pro Frame und Farbe (in der Spalte “name”) nur noch einen x-Wert habe.
Vielen vielen Dank für die Antwort!
Hi Remya,
kein Problem, wir können gerne auf Deutsch schreiben!
Ich habe ein Beispiel erstellt, das dir bei deiner Frage hoffentlich weiterhilft:
Viele Grüße,
Joachim
Hi Joachim,
how would you do this if you need to create a new variable, which shows per row which calculates this:
sepal.length for a specific row / mean per category
So for the first row, it would be like this: 5.1 / 5.006 (mean from setosa) = 1.018777. And for the 2nd row like this: 4.9/5.006 = 0.9788254 But then for every observation we have of course.
I can’t seem to figure it out. Thanks in advance
Hey Lang,
Thanks for the interesting question. It has inspired me to create a new tutorial on this topic. You can find it here.
I hope that helps!
Joachim
Hi Dear,
I am new to R. I am trying to calculate the mean output of the dynaCop model in R. The model is set to produce output per day, but yield_green is the yield of coffee beans, which come once a year. Therefore, this variable has zeros all year except when beans are harvested, so the mean value appears as 3.7 or 3.9, although the maximum value is 2028 (beans). I want to convert this variable into years and calculate the exact mean value. Because there are many other output variables in the model, the R environment refers to it as “Large list.”
Please Help me
Hey Faraz,
Could you please share your code and illustrate the structure of your data in some more detail?
Regards,
Joachim