Mode Imputation (How to Impute Categorical Variables Using R)
Mode imputation is easy to apply – but using it the wrong way might screw the quality of your data.
In the following article, I’m going to show you how and when to use mode imputation.
Before we can start, a short definition:
Mode imputation (or mode substitution) replaces missing values of a categorical variable by the mode of non-missing cases of that variable.
Impute with Mode in R (Programming Example)
Imputing missing data by mode is quite easy. For this example, I’m using the statistical programming language R (RStudio). However, mode imputation can be conducted in essentially all software packages such as Python, SAS, Stata, SPSS and so on…
Consider the following example variable (i.e. vector in R):
set.seed(951) # Set seed N <- 1000 # Number of observations vec <- round(runif(N, 0, 5)) # Create vector without missings vec_miss <- vec # Replicate vector vec_miss[rbinom(N, 1, 0.1) == 1] <- NA # Insert missing values table(vec_miss) # Count of each category # 0 1 2 3 4 5 # 86 183 207 170 174 90 sum(is.na(vec_miss)) # Count of NA values # 90
Our example vector consists of 1000 observations – 90 of them are NA (i.e. missing values).
Now lets substitute these missing values via mode imputation. First, we need to determine the mode of our data vector:
val <- unique(vec_miss[!is.na(vec_miss)]) # Values in vec_miss my_mode <- val[which.max(tabulate(match(vec_miss, val)))] # Mode of vec_miss
The mode of our variable is 2. With the following code, all missing values are replaced by 2 (i.e. the mode):
vec_imp <- vec_miss # Replicate vec_miss vec_imp[is.na(vec_imp)] <- my_mode # Impute by mode
That’s it. Imputation finished.
But do the imputed values introduce bias to our data? I’m going to check this in the following…
Did we Screw it up?
Did the imputation run down the quality of our data? The following graphic is answering this question:
missingness <- c(rep("No Missings", 6), rep("Post Imputation", 6)) # Pre/post imputation Category <- as.factor(rep(names(table(vec)), 2)) # Categories Count <- c(as.numeric(table(vec)), as.numeric(table(vec_imp))) # Count of categories data_barplot <- data.frame(missingness, Category, Count) # Combine data for plot ggplot(data_barplot, aes(Category, Count, fill = missingness)) + # Create plot geom_bar(stat = "identity", position = "dodge") + scale_fill_brewer(palette = "Set2") + theme(legend.title = element_blank())
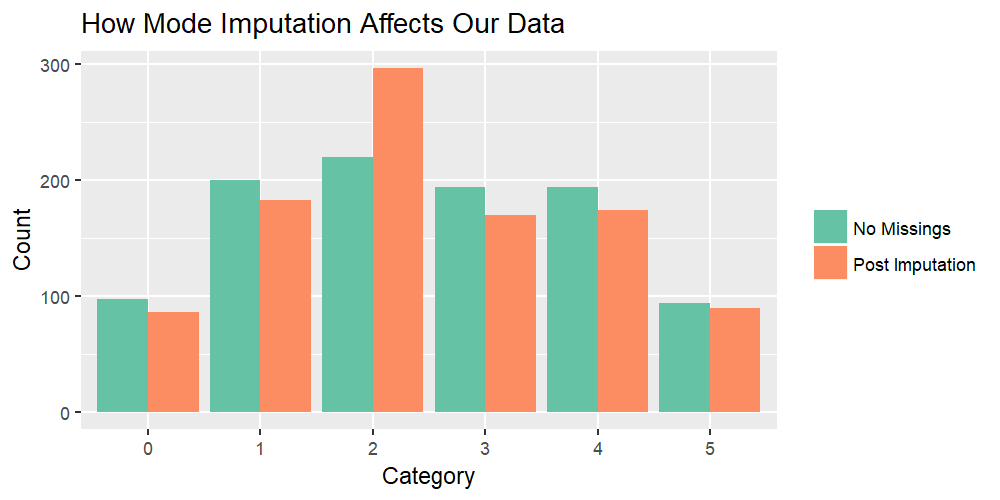
Graphic 1: Complete Example Vector (Before Insertion of Missings) vs. Imputed Vector
Graphic 1 reveals the issue of mode imputation:
The green bars reflect how our example vector was distributed before we inserted missing values. A perfect imputation method would reproduce the green bars.
However, after the application of mode imputation, the imputed vector (orange bars) differs a lot. While category 2 is highly over-represented, all other categories are underrepresented.
In other words: The distribution of our imputed data is highly biased!
Are There Better Alternatives?
As you have seen, mode imputation is usually not a good idea. The method should only be used, if you have strong theoretical arguments (similar to mean imputation in case of continuous variables).
You might say: OK, got it! But what should I do instead?!
Recent research literature advises two imputation methods for categorical variables:
- Multinomial logistic regression imputation
- Predictive mean matching imputation
Multinomial logistic regression imputation is the method of choice for categorical target variables – whenever it is computationally feasible. However, if you want to impute a variable with too many categories, it might be impossible to use the method (due to computational reasons). In this case, predictive mean matching imputation can help:
Predictive mean matching was originally designed for numerical variables. However, recent literature has shown that predictive mean matching also works well for categorical variables – especially when the categories are ordered (van Buuren & Groothuis-Oudshoorn, 2011). Even though predictive mean matching has to be used with care for categorical variables, it can be a good solution for computationally problematic imputations.
I would like to hear your opinion!
I’ve shown you how mode imputation works, why it is usually not the best method for imputing your data, and what alternatives you could use.
Now, I’d love to hear from your experiences!
Have you already imputed via mode yourself? Would you do it again?
Leave me a comment below and let me know about your thoughts (questions are very welcome)!
References
van Buuren, S., and Groothuis-Oudshoorn, C. G. (2011). MICE: Multivariate Imputation by Chained Equations in R. Journal of Statistical Software, 45(3).
Appendix
How to create the header graphic? There you go:
par(bg = "#1b98e0") # Background color par(mar = c(0, 0, 0, 0)) # Remove space around plot N <- 5000 # Sample size x <- round(runif(N, 1, 100)) # Uniform distribution x <- c(x, rep(60, 35)) # Add some values equal to 60 hist_save <- hist(x, breaks = 100) # Save histogram col <- cut(h$breaks, c(- Inf, 58, 59, Inf)) # Colors of histogram plot(hist_save, # Plot histogram col = c("#353436", "red", "#353436")[col], ylim = c(0, 110), main = "", xaxs="i", yaxs="i")
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.







22 Comments. Leave new
Mean and mode imputation may be used when there is strong theoretical justification. What can those justifications be? Can you please provide some examples.
Thank you very much for your well written blog on statistical concepts that are pre-digested down to suit students and those of us who are not statistician.
Cheers.
Hi Ugyen,
Thank you for your question and the nice compliment!
In practice, mean/mode imputation are almost never the best option. If you don’t know by design that the missing values are always equal to the mean/mode, you shouldn’t use it.
You may also have a look at this thread on Cross Validated to get more information on the topic.
Regards,
Joachim
Hi, thanks for your article. Can you provide any other published article for causing bias with replacing the mode in categorical missing values? Thanks
Hi Sara,
Thank you for the comment! For instance, have a look at Zhang 2016: “Imputations with mean, median and mode are simple but, like complete case analysis, can introduce bias on mean and deviation.”
Regards,
Joachim
Hi Joachim. What do you think about random sample imputation for categorical variables? Not randomly drawing from any old uniform or normal distribution, but drawing from the specific distribution of the categories in the variable itself.
As a simple example, consider the Gender variable with 100 observations. Male has 64 instances, Female has 16 instances and there are 20 missing instances. Before imputation, 80% of non-missing data are Male (64/80) and 20% of non-missing data are Female (16/80). After variable-specific random sample imputation (so drawing from the 80% Male 20% Female distribution), we could have maybe 80 Male instances and 20 Female instances.
Imputing this way by randomly sampling from the specific distribution of non-missing data results in very similar distributions before and after imputation. If mode imputation was used instead, there would be 84 Male and 16 Female instances. More biased towards the mode instead of preserving the original distribution.
My question is: is this a valid way of imputing categorical variables? What are its strengths and limitations?
Hi Ismail,
Thank you for you comment! The advantage of random sample imputation vs. mode imputation is (as you mentioned) that it preserves the univariate distribution of the imputed variable. However, there are two major drawbacks:
1) You are not accounting for systematic missingness. Assume that females are more likely to respond to your questionnaire. This would lead to a biased distribution of males/females (i.e. too many females). This is already a problem in your observed data. By imputing the missing values based on this biased distribution you are introducing even more bias. Have a look at the “response mechanisms” MCAR, MAR, and MNAR.
2) You are introducing bias to the multivariate distributions. For instance, assume that you have a data set with sports data and in the observed cases males are faster runners than females. If you are imputing the gender variable randomly, the correlation between gender and running speed in your imputed data will be zero and hence the overall correlation will be estimated too low.
For those reasons, I recommend to consider polytomous logistic regression. Have a look at the mice package of the R programming language and the mice() function. Within this function, you’d have to specify the method argument to be equal to “polyreg”.
I hope that helps!
Joachim
my mood function predicted values belonging to one class for classification data… what to do now
Hi Inzamam,
Could you tell me some more details about your data? How does the input data look like?
Regards,
Joachim
Thank you Joachim for this article. In my experiences, I have inputed by mod et mean. I do the remarque as it’stands one so.
Hey Alassane,
Thank you for your comment! 🙂
In most cases, I’d recommend using other methods such as predictive mean matching imputation and hot deck imputation. However, this depends strongly on your specific data.
Regards
Joachim
I use multiple imputation approach (via Mice package), using the “polyreg” option. I will take a keen interest into evaluating distribution of the imputation
Hey Sam,
Your approach is usually a good choice, and basically always better than a simple mode imputation.
Regards,
Joachim
Thank you for the illustration. Is there an efficient way for mode imputation while grouping by several other categorical variables in a dataframe? I’ve been trying to do so with dplyr but haven’t been able to make it work.
Hey Harry,
Please have a look at the example code below. This code can be improved a lot in terms of efficiency. However, it should give you a good basis:
I hope that helps!
Joachim
Hi Joachim,
How do you impute the mode of the categorical column containing missing cells already replaced by NA e.g. In a column of red and blue with the former being the most frequent, how do you impute red in this case to fill in the NA cells? Will be grateful if you could shed some light on this. Thank you.
Hey Lola,
Could you provide some example data or illustrate the structure of your data in some more detail? I’m afraid I don’t understand your question properly.
Regards,
Joachim
Hallo Joachim,
Thank you so much for explanation, now I am a big fan of your channel on youtibe and Statistics Globe. In fact I’ve learned hear alot and I am not exaggerating when I say that I’ve learned here especially in practice more than what I studied in some statistic seminars. Thank you so much 🙂 I have a small question: in my dataset I have some binary variables like sex (male, female) and social background (migrant or native), the observations are already coded as numbers like 1 for male and 2 for females and the same for social background. I checked the type of the variable with r typeof command and it says Integer. should I then recode them as factor and change values to Male, Female before using the Multinomial logistic regression imputation or the PMM imputation. In general I am using the PMM imputation for all variables in my dataset..
Thank you so much
Hey Mahmoud,
First of all: Thank you so much for the very kind feedback! It’s really great to hear that my tutorials are helpful to you! 🙂
I’ll partly copy/paste my response to your other question on YouTube, because I think it’s relevant to this question as well:
Usually, I would use predictive mean matching only for numeric variables and polynomial regression (method “polyreg” in the mice package) for categorical variables. If your categorical variables are ordered, you may also use predictive mean matching for them. However, it is an ongoing discussion whether predictive mean matching is preferable compared to polyreg. For unordered categorical variables or for binary variables, I would not use predictive mean matching, but other methods such as polyreg or hot deck imputation.
Your binary variables should be coded as factors instead of integers (it doesn’t matter if it’s 1/0 or male/female). If this is done correctly, the mice package should automatically choose a different method (i.e. “logreg”) than pmm for these variables.
I have created a reproducible example on how such an imputation could look like:
Above, you can see a multiply imputed data set. The variable x1 was imputed by logreg, and the variables x2 and x3 by pmm.
Regards,
Joachim
Very good article! Thank you!!
I wanted to ask, I am struggling with a health care related dataset with 14 numeric variables, with about 30% of them missing. I want to fill the missing data in order to use them into a clustering technique.
I have tried mice but the further I can go is the first step of creating the imputed datasets. After that I cannot pool my results as long as I have not a response variable. I have read about maximum likelihood imputation and I don’t know if this could be an option. Thank you in advance!
Hi Ann,
Thank you for the kind comment, glad you found it helpful!
I think your question is very interesting. Unfortunately, I do not have much experience with clustering techniques, so I might be the wrong person to ask.
Could you maybe post your question to the Statistics Globe Facebook group? Perhaps somebody there has a good answer on this: https://www.facebook.com/groups/statisticsglobe
Regards,
Joachim
Dear Joachim,
Thank you for your article.
I have some questions about missing data imputation.
1. There is missingness in my dataset, some due to drop out from the measurement, some due to other reasons (experimenter’s mistake or external interference, etc. I’d like to know whether we should include the dropout missingness in the missing data imputation. The percentage of drop out missingness is around 40%. The missing data percentage is around 15% when excluding the dropout missingness.
2. My current study is part of a broader longitudinal study with a sample size of 193 participants. To minimize bias in data distribution, I have included all behavioral variables with missingness from the entire study in the imputation process. After imputing the missing data from the entire longitudinal study, I applied these imputed variables to a subset of the data, that is, our current study. Is this manipulation is proper?
I look forward to your reply. Thank you so much for your help.
Best regards,
Hello,
Sorry for the late reply Sue. I was on a vacation. Do you still need help?
Best,
Cansu