Extract Beta Coefficients from Linear Regression Model in R (Example)
In this tutorial, I’ll illustrate how to get standardized regression coefficients (also called beta coefficients or beta weights) from a linear model in R.
The article contains this:
Let’s get started…
Introduction of Example Data
Consider the following example data:
set.seed(2344637) # Create example data x1 <- rnorm(100) x2 <- rnorm(100) + 0.25 * x1 x3 <- rnorm(100) + 0.5 * x1 - 0.3 * x2 y <- rnorm(100) + 0.15 * x1 + 0.4 * x2 - 0.1 * x3 data <- data.frame(x1, x2, x3, y) head(data) # Print head of example data
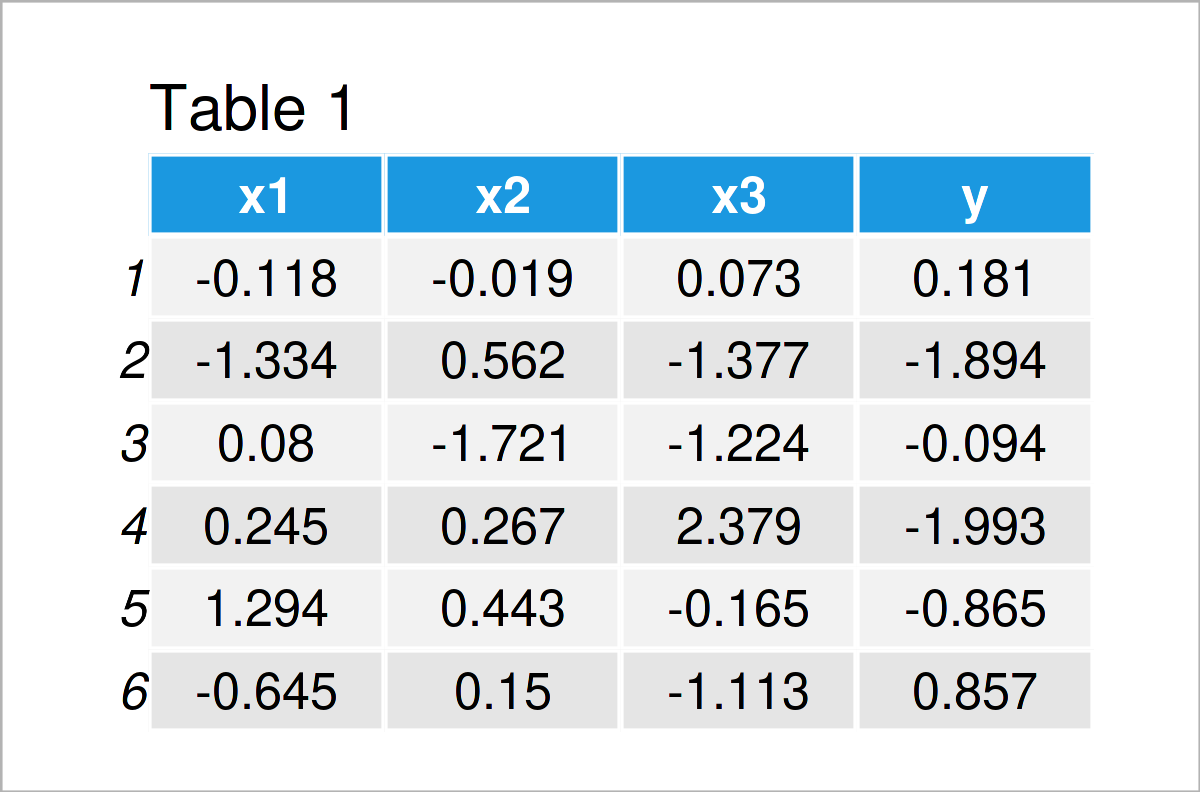
Table 1 shows that our example data contains four columns. The variables x1-x3 will be used as predictors and the variable y as target variable.
Next, we can estimate a linear regression model based on our data using the lm function:
my_mod <- lm(y ~ ., data) # Estimate linear regression model summary(my_mod) # Print summary statistics # Call: # lm(formula = y ~ ., data = data) # # Residuals: # Min 1Q Median 3Q Max # -2.52046 -0.72756 0.04412 0.73519 2.82633 # # Coefficients: # Estimate Std. Error t value Pr(>|t|) # (Intercept) 0.27485 0.10599 2.593 0.0110 * # x1 0.13312 0.12173 1.094 0.2769 # x2 0.22596 0.10180 2.220 0.0288 * # x3 -0.08231 0.11531 -0.714 0.4771 # --- # Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1 # # Residual standard error: 1.049 on 96 degrees of freedom # Multiple R-squared: 0.06315, Adjusted R-squared: 0.03388 # F-statistic: 2.157 on 3 and 96 DF, p-value: 0.09811
The previous output shows the summary statistics for our regression model.
However, this output does not show the beta coefficients. This is what we are going to compute next!
Note: the following two examples might produce slightly different results in case your data contains NA values. Please have a look at the comments of Thomas J. below this tutorial to read more on this topic.
Example 1: Extract Standardized Coefficients from Linear Regression Model Using Base R
In this example, I’ll explain how to calculate beta weights based on a linear regression model using the basic installation of the R programming language.
More precisely, we are using the lm, data.frame, and scale functions.
Consider the following R code and its output below:
lm(data.frame(scale(my_mod$model))) # Get standardized regression coefficients # Call: # lm(formula = data.frame(scale(my_mod$model))) # # Coefficients: # (Intercept) x1 x2 x3 # 5.070e-17 1.441e-01 2.200e-01 -9.387e-02
As you can see, we have returned the beta coefficients corresponding to our linear regression model.
Note that the code of this example was provided by Dr. R. H. Red Owl – thanks a lot to him! Please have a look at his comments below this tutorial to get more info on this code.
Example 2: Extract Standardized Coefficients from Linear Regression Model Using lm.beta Package
Alternatively to the functions of Base R (as explained in Example 1), we can also use the lm.beta package to get the beta coefficients.
In order to use the functions of the lm.beta package, we first have to install and load lm.beta to R:
install.packages("lm.beta") # Install lm.beta package library("lm.beta") # Load lm.beta package
In the next step, we can apply the lm.beta function to the lm model object that we have created before:
lm.beta(my_mod) # Get standardized regression coefficients # Call: # lm(formula = y ~ ., data = data) # # Standardized Coefficients:: # (Intercept) x1 x2 x3 # 0.0000000 0.1441121 0.2199585 -0.0938736
The previous output shows our standardized regression coefficients.
Video, Further Resources & Summary
Do you need further info on the R syntax of this article? Then I recommend watching the following video instruction on my YouTube channel. In the video, I’m explaining the R syntax of this article in R:
Additionally, you could have a look at some of the other articles on my website:
- Extract Fitted Values from Regression Model in R
- Exclude Specific Predictors from Linear Regression Model
- Extract Standard Error, t-Value & p-Value from Linear Regression Model
- Extract Residuals & Sigma from Linear Regression Model
- How to Extract the Intercept from a Linear Regression Model
- R Programming Overview
Summary: You have learned in this article how to calculate beta coefficients from a linear model in the R programming language. In case you have additional questions, please let me know in the comments section.







14 Comments. Leave new
Thanks for all your very helpful posts and Youtubes.
As you probably already know, R stores a pointer to the raw data in the regression model. That allows us to get the beta coefficients with a single line of Base R code after running a linear model.
Hey,
Thanks a lot for this very nice alternative code! I have added another example (see Example 1), which uses your code to produce the beta coefficients.
Regards,
Joachim
Thanks again for the additional syntax, this is great! 🙂
Regards,
Joachim
Hi, Joachim! What if I want to assign the beta and r value from the exponential regression model to a variable. How do I do it. THanks!
Hey Christian,
I assume this should be possible with the code of the present tutorial. Have you already tried this code?
Regards,
Joachim
Hi Joachim,
How can use the estimated coefficient of regression to predict another dataset?
Thanks.
Hi Christian,
Please have a look at my response to your other comment.
Regards,
Joachim
Hey Joachim,
Thank you very much for your help. I appreciate it.
Christian
You are very welcome, Christian.
Regards,
Joachim
Hi Joachim,
first of all many thanks for the great posts here and on Youtube. That always helps me a lot.
I noticed something with my own data set using the solutions shown here. The two examples work fine when the dataset is complete. However, if values are missing by chance, then you get slight differences.
In order to see the effect, one can first randomly remove a few values in e.g. column x1 and x3.
set.seed(16)
data[sample(1:nrow(data), 5), “x1”] <- NA
data[sample(1:nrow(data), 5), "x3"] <- NA
The beta coefficients can also be obtained via the following regression:
my_mod_standardized <- lm(scale(y) ~ scale(x1) + scale(x2) + scale(x3), data = data)
summary(my_mod_standardized)
If you now run the following code, you will notice slight differences:
my_mod <- lm(y ~ ., data)
lm(data.frame(scale(my_mod$model)))
lm.beta(my_mod)
You can also calculate the beta coefficients manually (see for the formula e.g. https://en.wikipedia.org/wiki/Standardized_coefficient). These correspond to the values of the regression.
my_mod$coefficients[2]*sd(data$x1, na.rm = TRUE)/sd(data$y, na.rm = TRUE) # beta coefficient of x1
my_mod$coefficients[3]*sd(data$x2, na.rm = TRUE)/sd(data$y, na.rm = TRUE) # beta coefficient of x2
my_mod$coefficients[4]*sd(data$x3, na.rm = TRUE)/sd(data$y, na.rm = TRUE) # beta coefficient of x3
Please let me know if I have missed anything.
Best regards,
Thomas
Hi Thomas,
Thanks a lot for the very kind words regarding my tutorials, glad you find them useful!
Regarding your question, I’ve just executed your code, and I’m also receiving different outputs. To be honest, I don’t know why this is happening. I assume that one of the functions must handle NA values differently than the other, but you would have to dive deeper into the help documentation of the different functions to read more about their handling of missing data.
Please let me know in case you find out why this happens, I’m also curious now.
Regards,
Joachim
Hi Joachim,
Thank you very much for your quick reply.
Now that I have also been gripped by curiosity, I have taken a closer look at the lm.beta() function. For the first possibility “Example 1: Extract Standardized Coefficients from Linear Regression Model Using Base R” I suspect the same reason that I think I have found now:
In your original data set, I had randomly removed 5 values in x1 and 5 values in x3. So there were a total of 10 rows with NA values.
With the lm.beta() function (https://rdrr.io/cran/lm.beta/src/R/lm.beta.R), all rows that contain an NA value in any variable are completely removed. This means that a data set with 90 rows is used for all calculations here. But this rather means that even for the standardisation of variables that actually had no NA value at all (x2 and y), only 90 observations are used.
In contrast, I calculated the standardisation according to the textbook formula (see e.g. Wooldridge, 2020, Introductory Econometrics, A Modern Approach, p. 184 f.), i.e. the standard deviations of x2 and y were calculated on the basis of the 100 existing observations.
This corresponds to the possibility using the regression method: lm(scale(y) ~ scale(x1) + scale(x2) + scale(x3), data = data).
As a sidenote, I noticed that the lm.beta() function does not standardise via the standard deviation, but via the root of the sum of the squared deviations (https://rdrr.io/cran/lm.beta/f/inst/doc/implementation.pdf). However, this is only generally valid if there are the same number of non-NA values in the respective variables.
I hope it helps you and all users here a little bit.
Best regards,
Thomas
Hi Thomas,
Once again, thank you very much for sharing this additional info. This will be very useful to other readers.
I have added a note in the tutorial that users should read your comments in case they have NA values in their data.
Thanks,
Joachim