Include All Two-Way Interactions into Model in R (2 Examples)
This tutorial demonstrates how to add all two-way interaction effects to a linear model in R programming.
The focus of this tutorial is linear models, like linear regression models, generalized linear regression models, linear mixed models, etc.
Non-linear models are out of the scope of the tutorial since the interaction terms may not be simple multiplicative terms like in linear models. Instead, they can be more complex expressions involving variables and model parameters.
The table of content looks as follows:
Let’s start right away!
Example Data
The first step is to generate the data to be used in the following examples. I created three independent variables, x1, x2, and x3, and two dependent variables, y_linear, and y_binary. Then I stored them in a dataset called my_data, as shown below.
set.seed(42) n <- 100 # Generate sample data x1 <- rnorm(n) x2 <- rnorm(n) x3 <- rnorm(n) y_linear <- 1 + 2 * x1 + 3 * x2 + x1 * x2 + rnorm(n, sd = 1) y_binary <- ifelse(1 + 2 * x1 - 3 * x2 + 0.5 * x3 + x1 * x2 - x1 * x3 + rnorm(n, sd = 1) > 0, 1, 0) my_data <- data.frame(x1, x2, x3, y_linear, y_binary) # Create my_data head(my_data)
Please be aware that the set.seed() was used to generate the same values at each run of the number generation. For further details, see Why & How to Set a Random Seed in R.
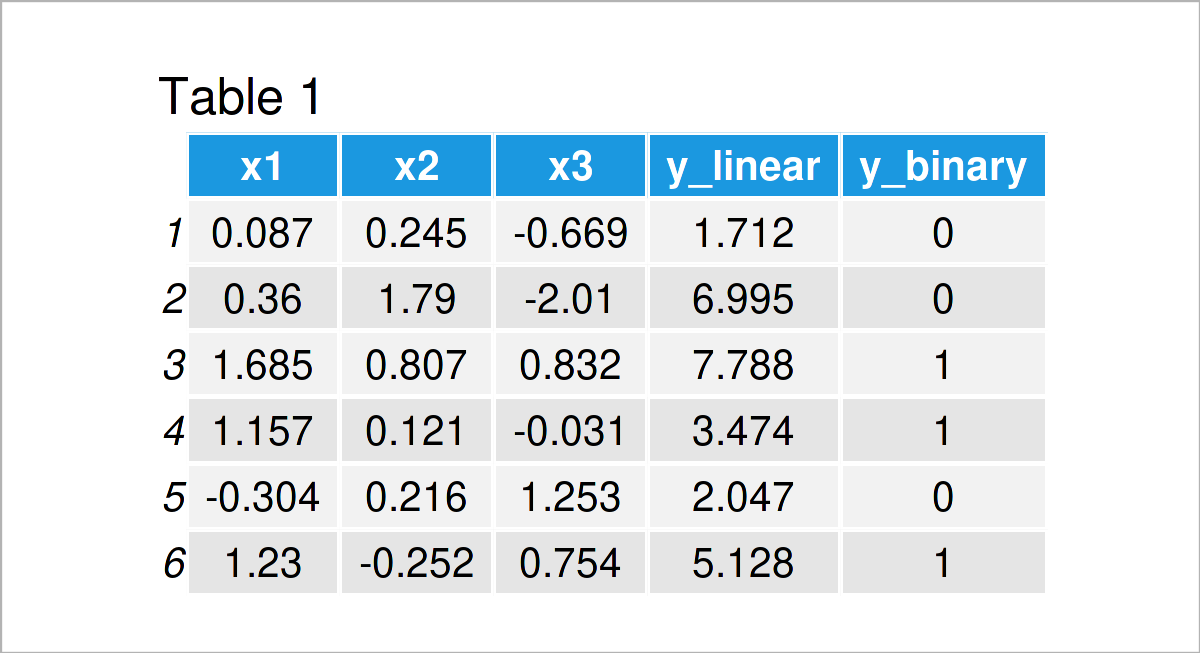
In Table 1, you can see that all variables have been successfully included in my_data.
Example 1: Set All Two-Way Interactions using ^2
One can add all two-way interactions using the multiplication operator * as usual, like x1*x2. However, it might be time- and attention-consuming if there are many variables to consider. In Example 1, I’ll illustrate how to use the exponent operator ^ as a shortcut to include all two-way interactions instead of adding all pairwise multiplications one by one. See below how it is implemented.
lm_model <- lm(y_linear ~ (x1 + x2 + x3)^2, data = my_data) # Fit lm model summary(lm_model) # Summary of results # Call: # lm(formula = y_linear ~ (x1 + x2 + x3)^2, data = my_data) # # Residuals: # Min 1Q Median 3Q Max # -2.1722 -0.5290 -0.1009 0.6757 2.1412 # # Coefficients: # Estimate Std. Error t value Pr(>|t|) # (Intercept) 1.28250 0.09634 13.312 <2e-16 *** # x1 2.03018 0.09388 21.625 <2e-16 *** # x2 3.08226 0.10683 28.853 <2e-16 *** # x3 0.04513 0.11361 0.397 0.692 # x1:x2 0.95330 0.09507 10.027 <2e-16 *** # x1:x3 -0.11813 0.10891 -1.085 0.281 # x2:x3 0.07627 0.11754 0.649 0.518 # --- # Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1 # # Residual standard error: 0.9504 on 93 degrees of freedom # Multiple R-squared: 0.9555, Adjusted R-squared: 0.9527 # F-statistic: 333.1 on 6 and 93 DF, p-value: < 2.2e-16
As seen, taking the square of (x1 + x2 + x3) helps the user include all two-way interactions x1:x2, x1:x3, and x2:x3 in addition to the main effects.
Example 2: Set All Two-Way Interactions using .^2
In this example, I’ll show a further shortcut to include all two-way interactions in a linear model. Let’s pick another type of model this time. I opted for logistic regression, which is computed by the generalized linear model function glm(). Relatedly, the dependent variable will be y_binary instead of y_linear.
Different from Example 1, I will use . instead of the formula of (x1 + x2 + x3). Let’s see if the command includes all two-way interactions successfully!
glm_model <- glm(y_binary ~ .^2 , data = my_data, family = binomial) # Fit glm model summary(glm_model) # Summary of results # Call: # glm(formula = y_binary ~ .^2, family = binomial, data = my_data) # # Deviance Residuals: # Min 1Q Median 3Q Max # -2.48645 -0.00848 0.00513 0.12757 2.18136 # # Coefficients: # Estimate Std. Error z value Pr(>|z|) # (Intercept) 0.69074 0.76331 0.905 0.36550 # x1 3.97820 1.60293 2.482 0.01307 * # x2 -7.39078 2.52667 -2.925 0.00344 ** # x3 1.37089 0.82534 1.661 0.09671 . # y_linear 0.06587 0.57745 0.114 0.90918 # x1:x2 1.20219 4.54634 0.264 0.79145 # x1:x3 -2.84938 1.69833 -1.678 0.09340 . # x1:y_linear 0.17037 0.70335 0.242 0.80861 # x2:x3 -1.61301 3.61481 -0.446 0.65544 # x2:y_linear 0.30464 0.44466 0.685 0.49327 # x3:y_linear 0.33274 0.66416 0.501 0.61638 # --- # Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1 # # (Dispersion parameter for binomial family taken to be 1) # # Null deviance: 128.207 on 99 degrees of freedom # Residual deviance: 30.737 on 89 degrees of freedom # AIC: 52.737 # # Number of Fisher Scoring iterations: 9
As seen, the output contains some undesired two-way interactions. The reason is that when . is used to refer to all independent variables, all variables in the dataset are treated as independent variables except for the dependent variable. So to say, using . is occasionally useful.
For the sake of demonstration, we can subset our dataset and rerun the code.
my_data_new<-subset(my_data, select=c(-y_linear)) # Subset data glm_model <- glm(y_binary ~ .^2 , data = my_data_new, family = binomial) # Fit glm model summary(glm_model) # Summary of results # Call: # glm(formula = y_binary ~ .^2, family = binomial, data = my_data_new) # # Deviance Residuals: # Min 1Q Median 3Q Max # -2.00030 -0.04383 0.00134 0.10373 2.70980 # # Coefficients: # Estimate Std. Error z value Pr(>|z|) # (Intercept) 1.88089 0.87514 2.149 0.03162 * # x1 5.02280 1.66940 3.009 0.00262 ** # x2 -7.69973 2.36412 -3.257 0.00113 ** # x3 0.03014 0.68016 0.044 0.96466 # x1:x2 1.98879 1.03212 1.927 0.05399 . # x1:x3 -2.57986 1.05850 -2.437 0.01480 * # x2:x3 2.57605 1.61511 1.595 0.11072 # --- # Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1 # # (Dispersion parameter for binomial family taken to be 1) # # Null deviance: 131.791 on 99 degrees of freedom # Residual deviance: 27.009 on 93 degrees of freedom # AIC: 41.009 # # Number of Fisher Scoring iterations: 9
As seen, all two-way interactions have been successfully appended to the model. You can prefer the method most convenient for your case.
Video, Further Resources & Summary
Do you need more information on the examples in this article? Then I recommend watching the following video on my YouTube channel. In the video tutorial, I’m explaining the R code of this tutorial in RStudio.
The YouTube video will be added soon.
In addition, you might want to have a look at the related tutorials on this homepage:
- Exclude Specific Predictors from Linear Regression Model in R
- Extract Beta Coefficients from Linear Regression Model in R
- Extract Fitted Values from Regression Model in R
- Remove Intercept from Regression Model in R
- The R Programming Language
Summary: At this point of the tutorial, you should know how to compute all the two-way interaction effects in a linear model in R programming. In the comments section below, let me know if you have any additional questions.
This page was created in collaboration with Cansu Kebabci. Look at Cansu’s author page to get more information about her professional background, a list of all his tutorials, as well as an overview of her other tasks on Statistics Globe.







