R ggplot2 Error: Aesthetics must be either length 1 or the same as the data
In this tutorial you’ll learn how to fix the error message “Aesthetics must be either length 1 or the same as the data” in the R programming language.
Table of contents:
Let’s start right away…
Example Data, Add-On Packages & Default Graph
As a first step, we’ll need to create some data that we can use in the examples later on:
data <- data.frame(x = LETTERS[1:5], # Create example data y = 1:5) data # Print example data # x y # 1 A 1 # 2 B 2 # 3 C 3 # 4 D 4 # 5 E 5
In case we want to draw our data with the ggplot2 package, we also have to install and load ggplot2 to R:
install.packages("ggplot2") # Install & load ggplot2 package library("ggplot2")
Example 1: Reproducing the Error: Aesthetics must be either length 1 or the same as the data
In Example 1, I’ll show how to replicate the error message “Aesthetics must be either length 1 or the same as the data” in R.
Have a look at the following R code:
ggplot(data, aes(x, y, fill = c("red", "blue"))) + # Try to draw ggplot2 plot geom_bar(stat = "identity") # Error: Aesthetics must be either length 1 or the same as the data (5): fill
As you can see, the RStudio console returns the error “”Aesthetics must be either length 1 or the same as the data”.
The reason for this is that we have specified the fill argument within the aes function to be equal a vector of length 2 (i.e. c(“red”, “blue”)).
However, our example data has five categories and therefore doesn’t know which filling color should be used for which category.
Let’s solve this problem!
Example 2: Fixing the Error: Aesthetics must be either length 1 or the same as the data
This example shows how to deal with the ggplot2 error “Aesthetics must be either length 1 or the same as the data”.
As discussed before, we have to specify a filling color for each group in our data. The easiest way to do this is that we set fill to be equal to our grouping variable (i.e. x):
ggplot(data, aes(x, y, fill = x)) + # Properly drawing ggplot2 plot geom_bar(stat = "identity")
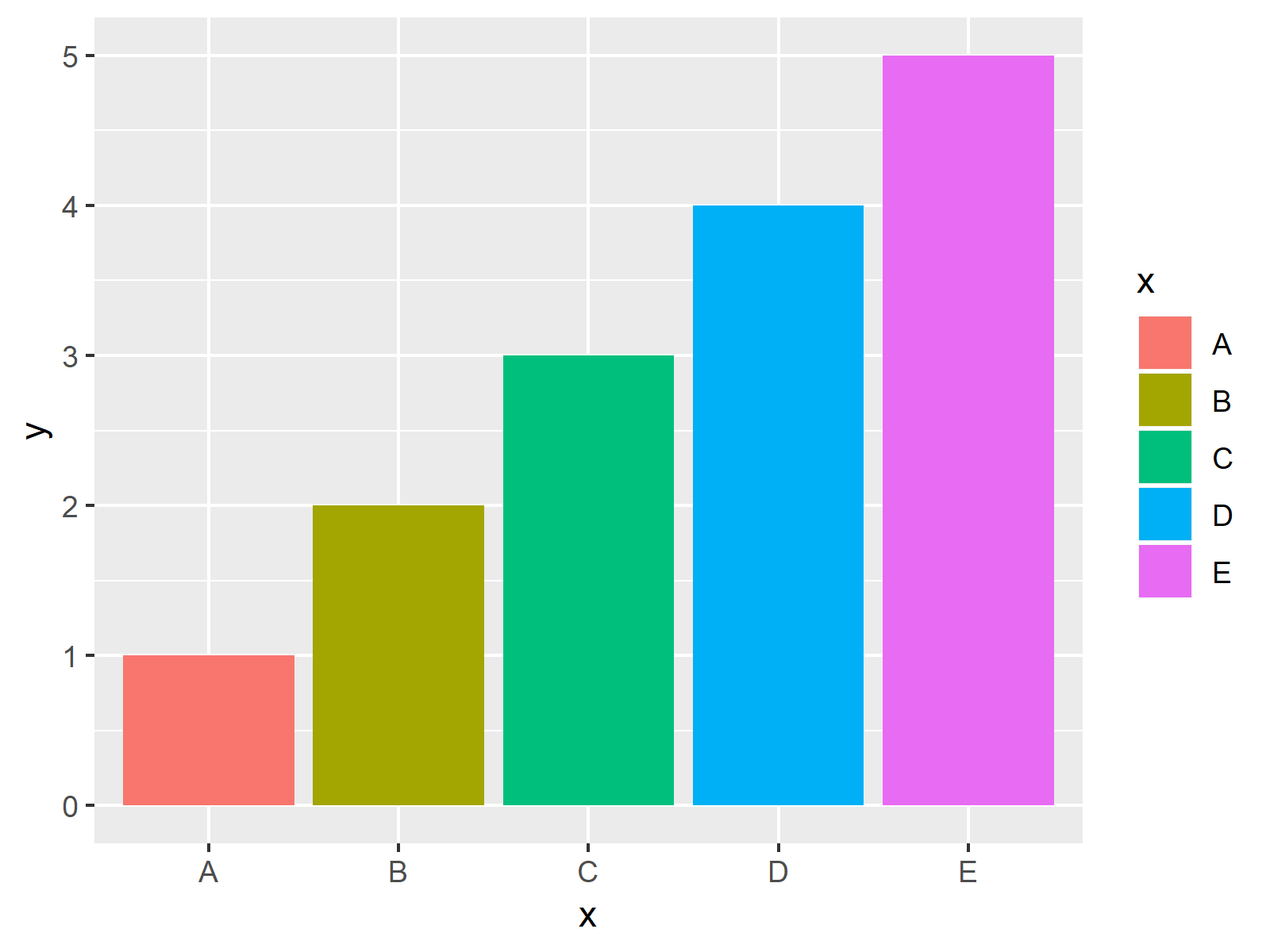
As illustrated in Figure 1, the previous syntax created a ggplot2 barplot in R. Looks good!
Video & Further Resources
I have recently released a video on my YouTube channel, which shows the R programming syntax of this tutorial. Please find the video below:
Furthermore, I can recommend to have a look at the other articles of my homepage.
- ggplot2 Error: stat_count() must not be used with a y aesthetic
- Error – Undefined Columns Selected when Subsetting Data Frame
- Dealing with Warnings & Errors in R (Cheat Sheet)
- R Programming Language
Summary: You learned in this article how to handle the error “Aesthetics must be either length 1 or the same as the data” in R programming. In case you have any further questions, let me know in the comments.
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.







14 Comments. Leave new
Hi,
I was trying to do the following code, and it hasn’t worked out.
Could you please help me know, where I went wrong:
I have picked the inbuilt set mpg and also run the suitable packages.
I get the error: Error: Aesthetics must be either length 1 or the same as the data (234): x, y and fill
set.seed(10)
names<-c(rep("A",20),
rep("B",5),
rep("C",30),
rep("D",100))
value<- c(sample(2:5, 20, replace=T),
sample(4:10, 5, replace=T),
sample(1:7, 30, replace=T),
sample(3:8, 100 ,replace=T))
data1<-data.frame(names,value)
data1
ggplot(data=mpg,(aes(x=names,y=value,fill=names))) +
geom_boxplot( stat="identity")
Thanks and kind regards,
Ananya
Hey Ananya,
Your code works for me when I do the following changes:
My final code looks like this:
And the resulting plot looks like this:
Regards,
Joachim
CHD <- rawdata$anychd
CHD2 <- as.character(CHD)
prev_CHD <- factor(CHD2, levels = c(0,1), labels = c("NO", "YES"))
age <- rawdata$AGE
ggplot(rawdata, aes(x=prev_CHD, y=age, coll = prev_CHD)) + geom_boxplot() + theme(legend.position = "none") + xlab("CHD Event") +ylab("Age (years)") + ggtitle("Relationship Between Age & Presence of CHD Event")
I also get the same error and was wondeirng why? Thanks!
Hey Charlotte,
You should store all the relevant information for the plot within a single data frame, not in separate data objects.
You may create a data frame containing prev_CHD and age and run the code again based on this data frame.
Regards,
Joachim
Hello,
I am just trying to generate a GSEA plot via plotEnrichment(). I did the following:
plotEnrichment(examplePathways[[“5991130_Programmed_Cell_Death”]],
exampleRanks) + labs(title=”Programmed Cell Death”)
And I received this message:
Error in `check_aesthetics()`:
! Aesthetics must be either length 1 or the same as the data (1): x and xend
Run `rlang::last_error()` to see where the error occurred.
I am not sure how to resolve this. Please help. Thanks
Hey Mike,
I’m sorry for the delayed response, I’ve been on vacation for the last couple of days. Do you still need help with this error message?
Regards,
Joachim
Hi, I am trying to produce a forest plot for my regression analysis using the following code:
plot1 = ggplot(output6_0b[[8]] %>%
mutate(ci.lb = ifelse(ci.lb cutoff_upper,
cutoff_upper, ci.ub)) ) +
geom_point(aes(y = name, x = estimate),
shape = 18, size = 5) +
geom_errorbarh(aes(xmin = ci.lb, xmax = ci.ub,
y = name), height = 0.25) +
geom_vline(xintercept = 1, color = “red”,
linetype = “dashed”, cex = 1, alpha = 0.5) +
# scale_x_continuous(limit = c(1.01*cutoff_lower, 1.01*cutoff_upper) ) +
xlab(“Lnrr”) +
ylab(“”) +
theme_bw() +
theme(panel.border = element_blank(),
panel.background = element_blank(),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
axis.line = element_line(colour = “black”),
axis.text.y = element_text(size = 12, colour = “black”),
axis.text.x.bottom = element_text(size = 12, colour = “black”),
axis.title.x = element_text(size = 12, colour = “black”))
table_base <- ggplot(output6_0b[[8]], aes(y=name)) +
ylab(NULL) + xlab(" ") +
theme(plot.title = element_text(hjust = 0.5, size=12),
axis.text.x = element_text(color="white", hjust = -3, size = 25), ## This is used to help with alignment
axis.line = element_blank(),
axis.text.y = element_blank(),
axis.ticks = element_blank(),
axis.title.y = element_blank(),
legend.position = "none",
panel.background = element_blank(),
panel.border = element_blank(),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
plot.background = element_blank())
## OR point estimate table
tab1 <- table_base +
labs(title = "space") +
geom_text(aes(y = rev(name), x = 1, label = sprintf("%0.1f", round(estimate, digits = 1))), size = 4) + ## decimal places
ggtitle("lnrr")
## 95% CI table
tab2 <- table_base +
geom_text(aes(y = rev(name), x = 4, label = CI), size = 4) +
ggtitle("95% CI")
lay <- matrix(c(1,1,1,1,1,1,1,1,1,1,2,3,3), nrow = 1)
grid.arrange(plot1, tab1, tab2, layout_matrix = lay)
but when I run the last line of the code I get the same error. I tried to find out the error but unfortunately could not. Could you please suggest why this error might appear and how to deal with it in this case? Thank you
Hi Milana,
This is difficult to tell without seeing your data. I assume output6_0b[[8]] is a data.frame object? Could you please share the output when you run the following code?
Regards,
Joachim
Hi Joachim,
Here they are
> head(output6_0b[[8]])
# A tibble: 4 × 7
name estimate se zval pval ci.lb ci.ub
1 intrcpt 2.98 2.10 1.42 0.156 -1.13 7.09
2 duration_exp -0.0387 0.0226 -1.71 0.0868 -0.0830 0.00559
3 application_rate -0.00554 0.0140 -0.395 0.693 -0.0330 0.0220
4 fertilizer_app_rate 0.000688 0.000911 0.755 0.450 -0.00110 0.00247
> str(output6_0b[[8]])
tibble [4 × 7] (S3: tbl_df/tbl/data.frame)
$ name : chr [1:4] “intrcpt” “duration_exp” “application_rate” “fertilizer_app_rate”
$ estimate: num [1:4] 2.980461 -0.038696 -0.005538 0.000688
$ se : num [1:4] 2.098382 0.022598 0.014032 0.000911
$ zval : num [1:4] 1.42 -1.712 -0.395 0.755
$ pval : num [1:4] 0.1555 0.0868 0.6931 0.4502
$ ci.lb : num [1:4] -1.1323 -0.083 -0.033 -0.0011
$ ci.ub : num [1:4] 7.09321 0.00559 0.02196 0.00247
Regards,
Milana
Hey Milana,
Thank you for the additional info.
I have tried to reproduce your data as shown below:
However, when I now run the first lines of your code, I get an error due to the data object cutoff_upper. Where is this data object coming from?
Furthermore, it seems like there is something wrong with the specification of the ifelse function: ifelse(ci.lb cutoff_upper, cutoff_upper, ci.ub) Why is there a space between ci.lb & cutoff_upper?
Please try to make your question reproducible.
Regards,
Joachim
Hi, could you please check where I made error. I am trying to created graph with forecast.
My code:
ggplot(data=myxts_final,aes (x=index(myxts_final)))+
ggtitle(“FTSE Close Index Forecasts”)+
geom_line(aes(y=dFT_xts2, color=”Spread”), linetype=”solid”, color=”black”)+
geom_ribbon(aes(ymin = y.lo80, ymax = y.hi80),fill = “blue”, alpha = 0.2)+
geom_ribbon(aes(ymin = y.lo95, ymax = y.hi95),fill = “blue”, alpha = 0.2)+
theme(axis.title.x=element_blank(),
axis.title.y=element_blank(),
axis.text.x=element_text(color=”black”,size=12),
axis.text.y=element_text(color=”black”, size=12),
panel.background=element_rect(fill=”white”),
panel.grid.minor=element_blank(),
panel.grid.major=element_blank(),
axis.line=element_line(color=”black”, size=1),
plot.title=element_text(face=”bold”))+ggeasy::easy_center_title()
The error I get:
Don’t know how to automatically pick scale for object of type .
Defaulting to continuous.
Error in `geom_line()`:
! Problem while computing aesthetics.
ℹ Error occurred in the 1st layer.
Caused by error in `check_aesthetics()`:
! Aesthetics must be either length 1 or the same as the data (6613)
✖ Fix the following mappings: `y`
Thank you!
Hello Alisha,
It is hard to know why this error is happening. But it looks like it is something to do with your y values. It says it should be continuous, but apparently, it is not so in your case. Please check your data type via the class() or typeof() functions. The error about the aesthetic length may also be related to the data type being wrong.
Regards,
Cansu
Hi! I keep coming across an error that says Error in `geom_point()`:
! Problem while computing aesthetics.
ℹ Error occurred in the 1st layer.
Caused by error:
! object ‘Concentration’ not found
Run `rlang::last_error()` to see where the error occurred.
I am not sure what to do!!1
This is around the time when my r code stops working:
#Plot the data and add best fit line
ggplot(my.data, aes(x=Concentration, y=Absorbance))+
geom_point()+
geom_smooth(method=’lm’, se=FALSE,
formula= y ~ 0 + x, color=”black”, linewidth=0.5)+
xlab(“Concentration (%)”)+
ylab(“Absorbance (OD)”)
Hello Katie,
The error message you are encountering indicates that the geom_point() function within ggplot is having trouble finding the Concentration object to use for plotting. Here are some possible reasons for this error and steps to troubleshoot it:
Column Name Typo: Ensure that the column name in your dataframe my.data exactly matches Concentration. R is case-sensitive, so make sure that the capitalization is identical.
Data Frame Attachment: Sometimes, if you have used attach(my.data), it can cause conflicts with ggplot. Make sure you haven’t attached the dataframe or try detaching it with detach(my.data) before running your ggplot code.
Correct Data Frame: Verify that my.data is the correct data frame and that it contains the column Concentration.
Check Data: Use str(my.data) to inspect the structure of your data frame. Make sure that the Concentration column exists and is in the correct format (typically numeric for a concentration value).
I hope one of these checks help to detect the cause of error.
Best,
Cansu