R Error in colMeans(x, na.rm = TRUE) : ‘x’ must be numeric (3 Examples)
This tutorial shows how to debug the “Error in colMeans(x, na.rm = TRUE) : ‘x’ must be numeric” in the R programming language.
The post is structured as follows:
Let’s dive into it:
Creating Example Data
Let’s first construct some exemplifying data:
set.seed(67932) # Create example data frame data <- data.frame(x1 = sample(LETTERS[1:3], 10, replace = TRUE), x2 = round(rnorm(10), 2), x3 = round(runif(10), 2)) data # Print example data frame
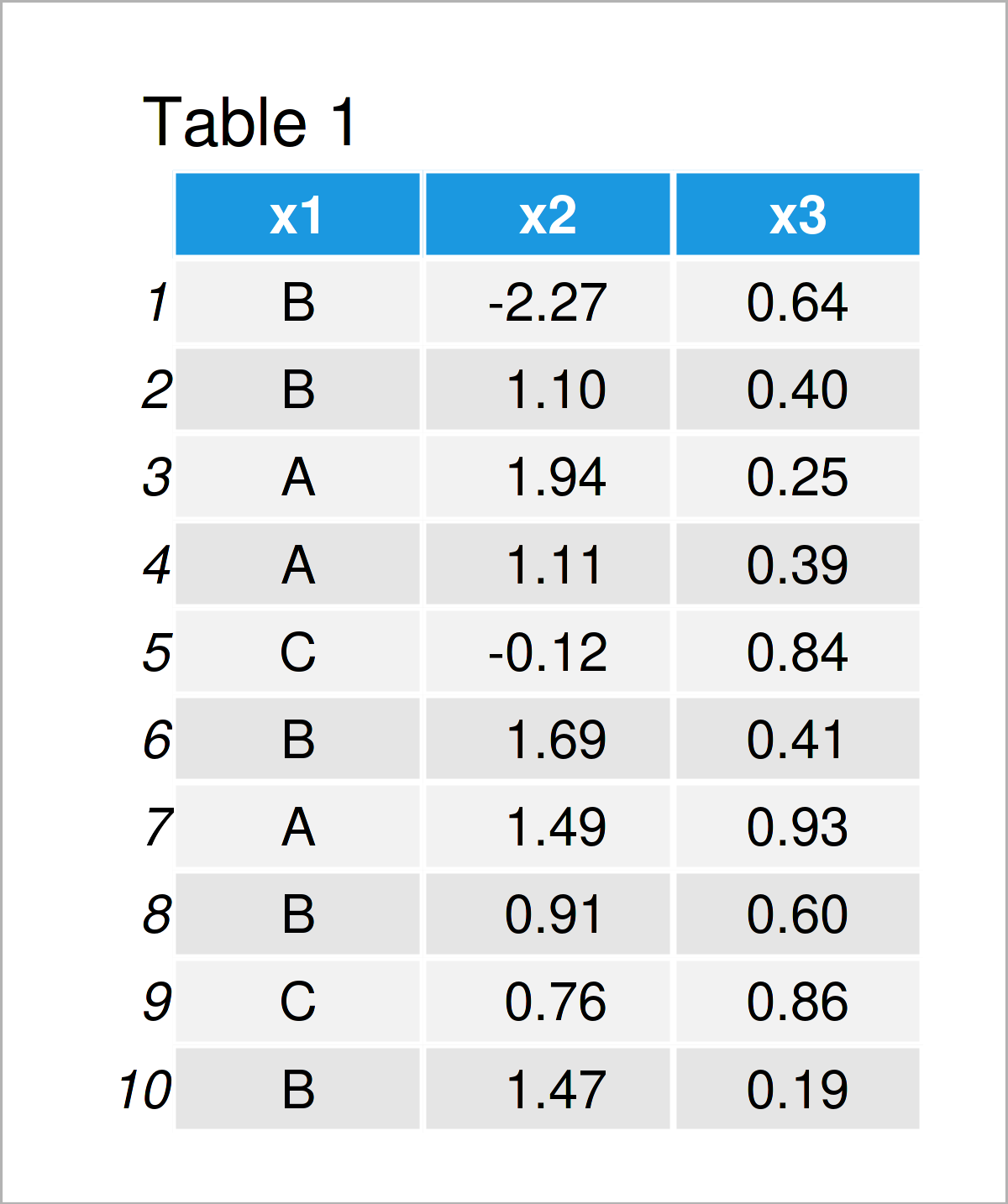
As you can see based on Table 1, our example data is a data frame and contains ten rows and three columns.
Example 1: Reproduce the Error in colMeans(x, na.rm = TRUE) : ‘x’ must be numeric
Example 1 explains how to replicate the “Error in colMeans(x, na.rm = TRUE) : ‘x’ must be numeric”.
Let’s assume that we want to apply a Principal Component Analysis based on these data.
Then, we might try to apply the prcomp function to our data as shown below:
prcomp(data) # Try to apply prcomp function # Error in colMeans(x, na.rm = TRUE) : 'x' must be numeric
Unfortunately, the “Error in colMeans(x, na.rm = TRUE) : ‘x’ must be numeric” is returned.
The reason for this error message is that our data frame contains the column x1 which has the character class (the same error would appear in case of factor columns).
So how could we fix that? There are basically two alternatives, and I’m going to explain these alternatives in the following examples.
Keep on reading!
Example 2: Fix the Error by Removing Non-Numeric Columns
In this example, I’ll demonstrate how to drop all non-numeric variables from a data frame to avoid the “Error in colMeans(x, na.rm = TRUE) : ‘x’ must be numeric”.
We can use the unlist, lapply, and is.numeric functions to create such a data frame subset:
data_new1 <- data[ , unlist(lapply(data, # Remove non-numeric columns is.numeric))] data_new1 # Print updated data frame
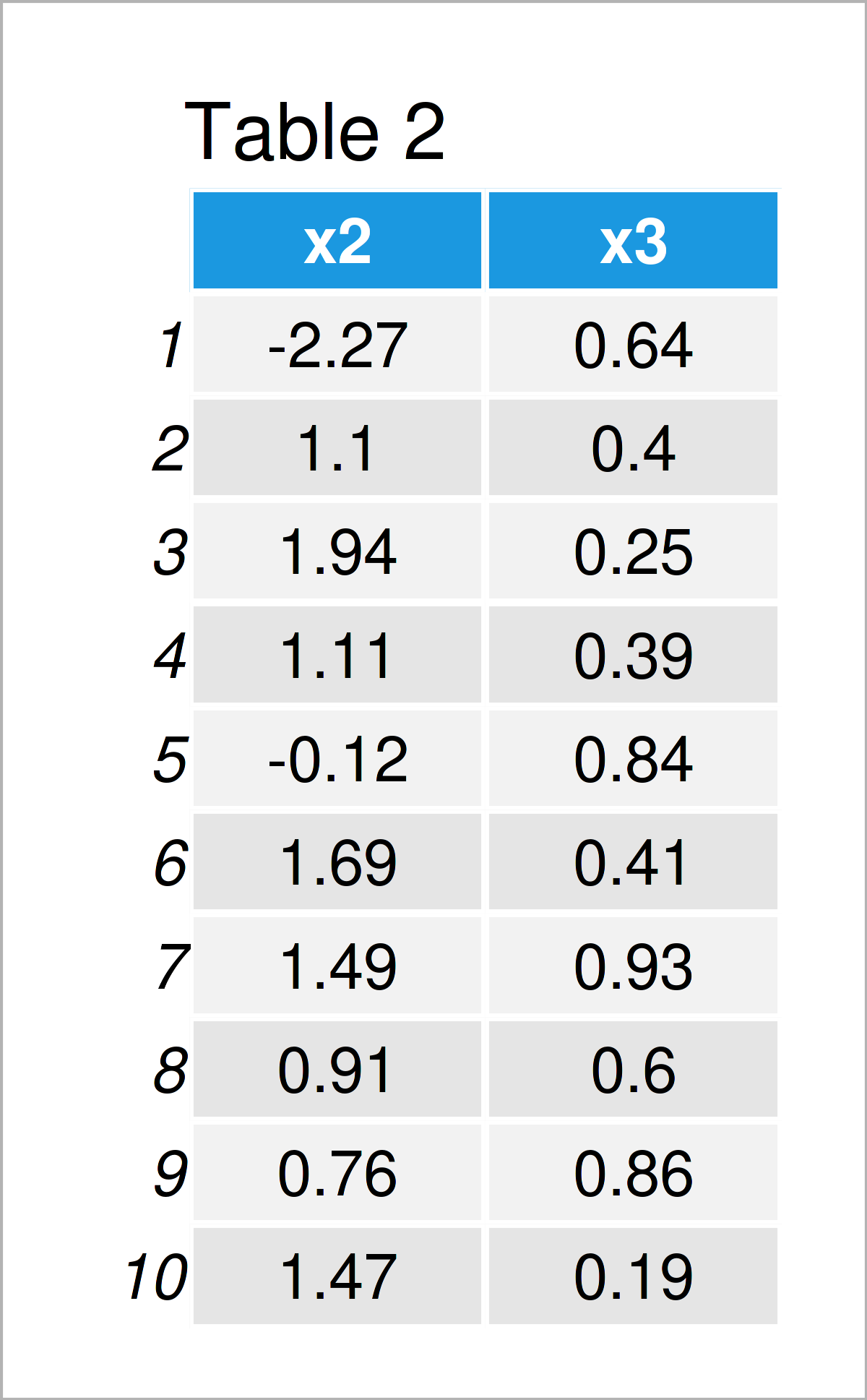
As shown in Table 2, we have created a new data frame with the previous R syntax. This data frame contains only the two numeric columns x2 and x3.
Next, we can apply the prcomp function to these data:
prcomp(data_new1) # Apply prcomp function # Standard deviations (1, .., p=2): # [1] 1.2283189 0.2428404 # # Rotation (n x k) = (2 x 2): # PC1 PC2 # x2 0.99647810 0.08385344 # x3 -0.08385344 0.99647810
Works fine!
Example 3: Fix the Error by Converting Non-Numeric Columns to Numbers
Example 3 demonstrates how to convert non-numeric categorical data to numeric data in order to get rid of the “Error in colMeans(x, na.rm = TRUE) : ‘x’ must be numeric”.
To accomplish this, we have to apply the as.numeric and as.factor functions to our non-numeric data frame column x1:
data_new2 <- data # Duplicate data frame data_new2$x1 <- as.numeric(as.factor(data_new2$x1)) # Convert categories to numbers data_new2 # Print updated data frame
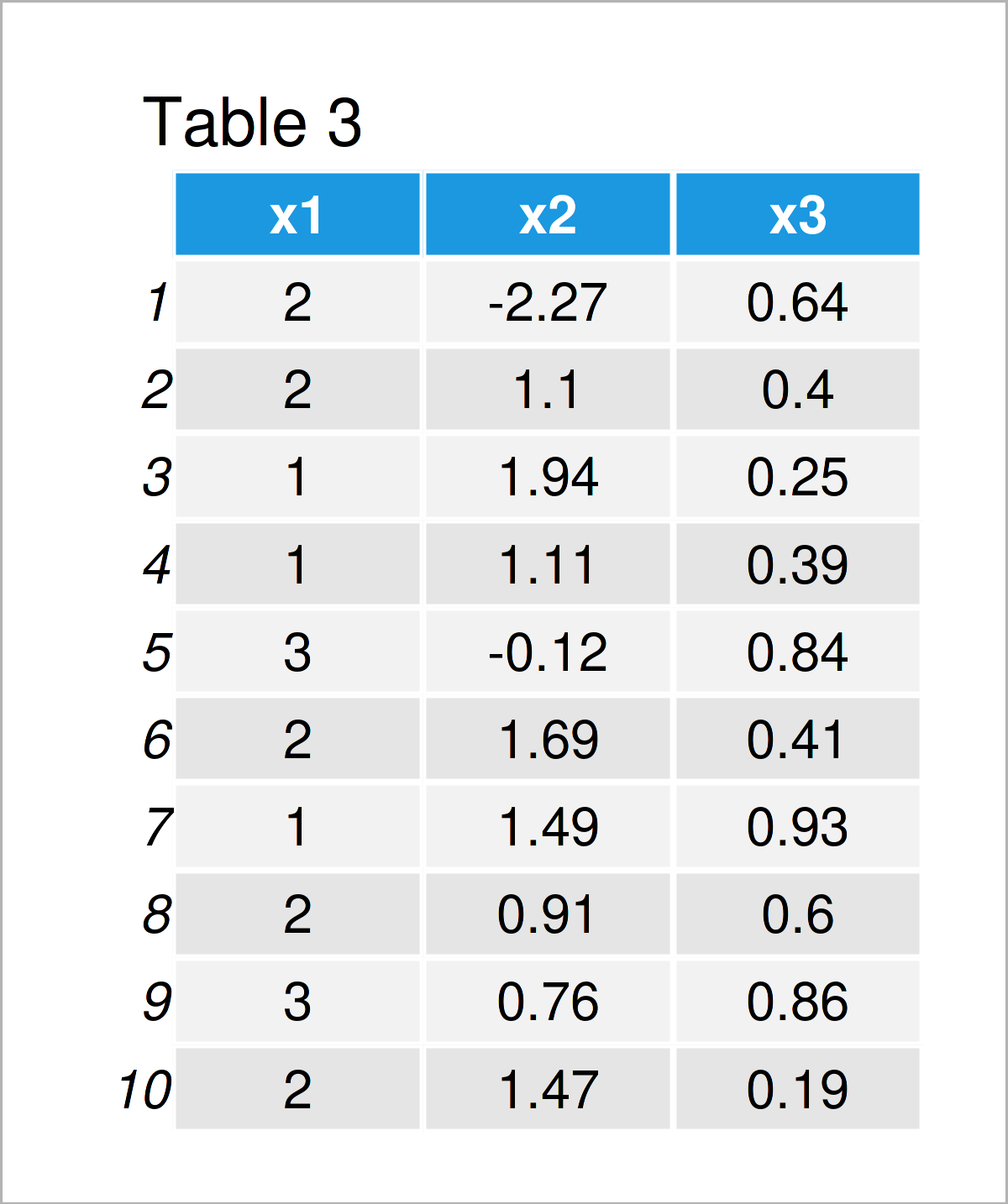
Table 3 shows the output of the previous code: We have transformed the categorical variable x1 into numbers.
Now, we can apply the prcomp function without any problems:
prcomp(data_new2) # Apply prcomp function # Standard deviations (1, .., p=3): # [1] 1.2734878 0.6608866 0.2316053 # # Rotation (n x k) = (3 x 3): # PC1 PC2 PC3 # x1 -0.3082818 0.9444851 -0.11362327 # x2 0.9471298 0.3158997 0.05614757 # x3 -0.0889241 0.0903067 0.99193609
Video & Further Resources
Have a look at the following video on my YouTube channel. In the video, I’m showing the content of this tutorial.
Furthermore, you may read the related tutorials on my website. A selection of articles can be found here.
- Error : ‘names’ attribute must be the same length as the vector
- Error in as.Date.numeric(X) : ‘origin’ must be supplied
- Error in sort.int(x, na.last, decreasing, …) : ‘x’ must be atomic
- Error in rowSums & colSums – ‘x’ must be an array of at least two dimensions
- Error in hist.default : ‘x’ must be numeric
- Solving Errors & Warnings in R
- Introduction to R Programming
Summary: You have learned in this article how to avoid the “Error in colMeans(x, na.rm = TRUE) : ‘x’ must be numeric” in R programming. Let me know in the comments section below, in case you have further questions.
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.






