Import & Merge Multiple CSV Files in R (2 Examples)
In this article, I’ll show you how to import and merge CSV files in the R programming language.
The page will contain the following topics:
Let’s do this!
Exemplifying Data
Before we can start with the examples, we need to create an exemplifying directory including multiple CSV files. First, we need to create several data frames…
data1 <- data.frame(id = 1:6, # Create first example data frame x1 = c(5, 1, 4, 9, 1, 2), x2 = c("A", "Y", "G", "F", "G", "Y")) data2 <- data.frame(id = 4:9, # Create second example data frame y1 = c(3, 3, 4, 1, 2, 9), y2 = c("a", "x", "a", "x", "a", "x")) data3 <- data.frame(id = 5:6, # Create third example data frame z1 = c(3, 2), z2 = c("K", "b"))
…and then we need to export these data frames as CSV files to our computer:
write.csv(data1, "C:/Users/Joach/Desktop/my_folder/data1.csv", # Write first example data row.names = FALSE) write.csv(data2, "C:/Users/Joach/Desktop/my_folder/data2.csv", # Write second example data row.names = FALSE) write.csv(data3, "C:/Users/Joach/Desktop/my_folder/data3.csv", # Write third example data row.names = FALSE)
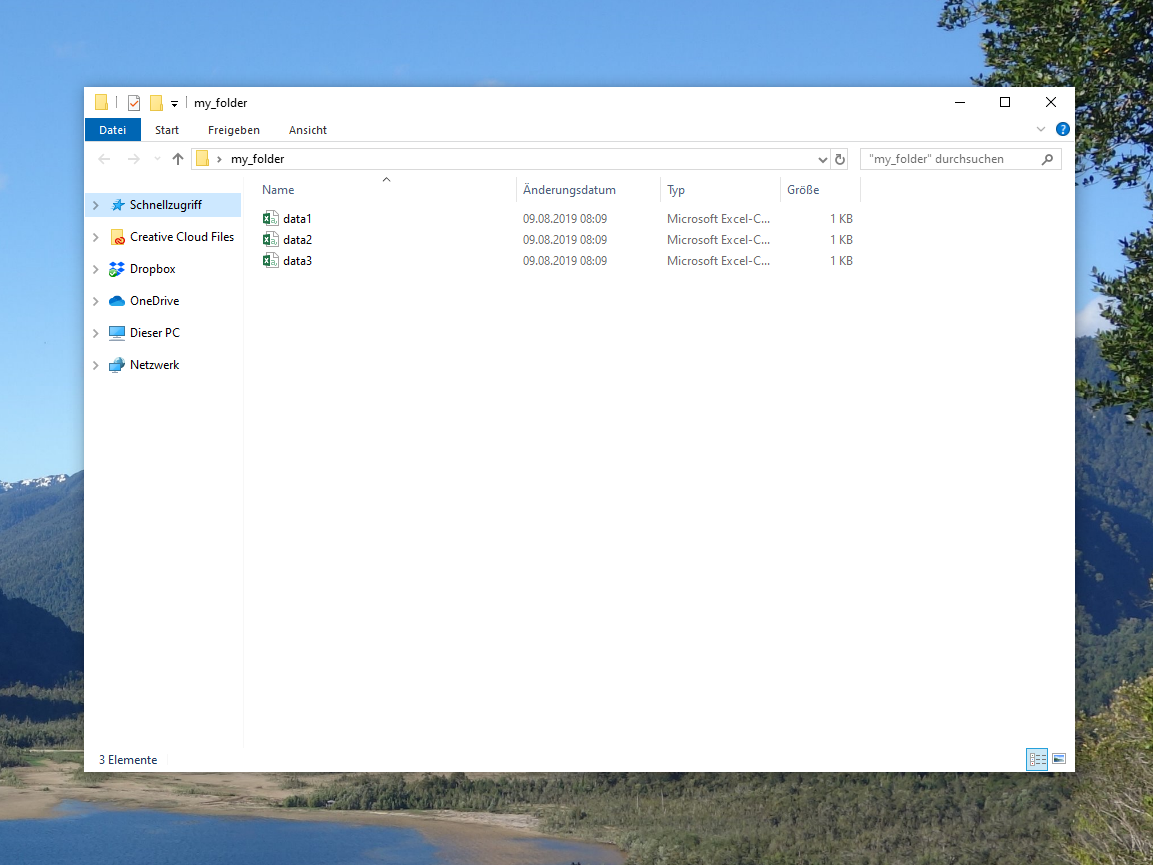
Figure 1: Exemplifying Directory with CSV Files.
Figure 1 illustrates how our example directory looks like. Now let’s import and combine these data sets in RStudio…
Example 1: Import & Row-Bind CSV Files in R
We need three R add-on packages for the following R syntax: dplyr, plyr, and readr. Let’s install and load these packages to R.
install.packages("dplyr") # Install dplyr package install.packages("plyr") # Install plyr package install.packages("readr") # Install readr package library("dplyr") # Load dplyr package library("plyr") # Load plyr package library("readr") # Load readr package
Now, we can import and merge the example CSV files based on the list.files, lapply, read_csv, and bind_rows functions:
data_all <- list.files(path = "C:/Users/Joach/Desktop/my_folder", # Identify all CSV files pattern = "*.csv", full.names = TRUE) %>% lapply(read_csv) %>% # Store all files in list bind_rows # Combine data sets into one data set data_all # Print data to RStudio console
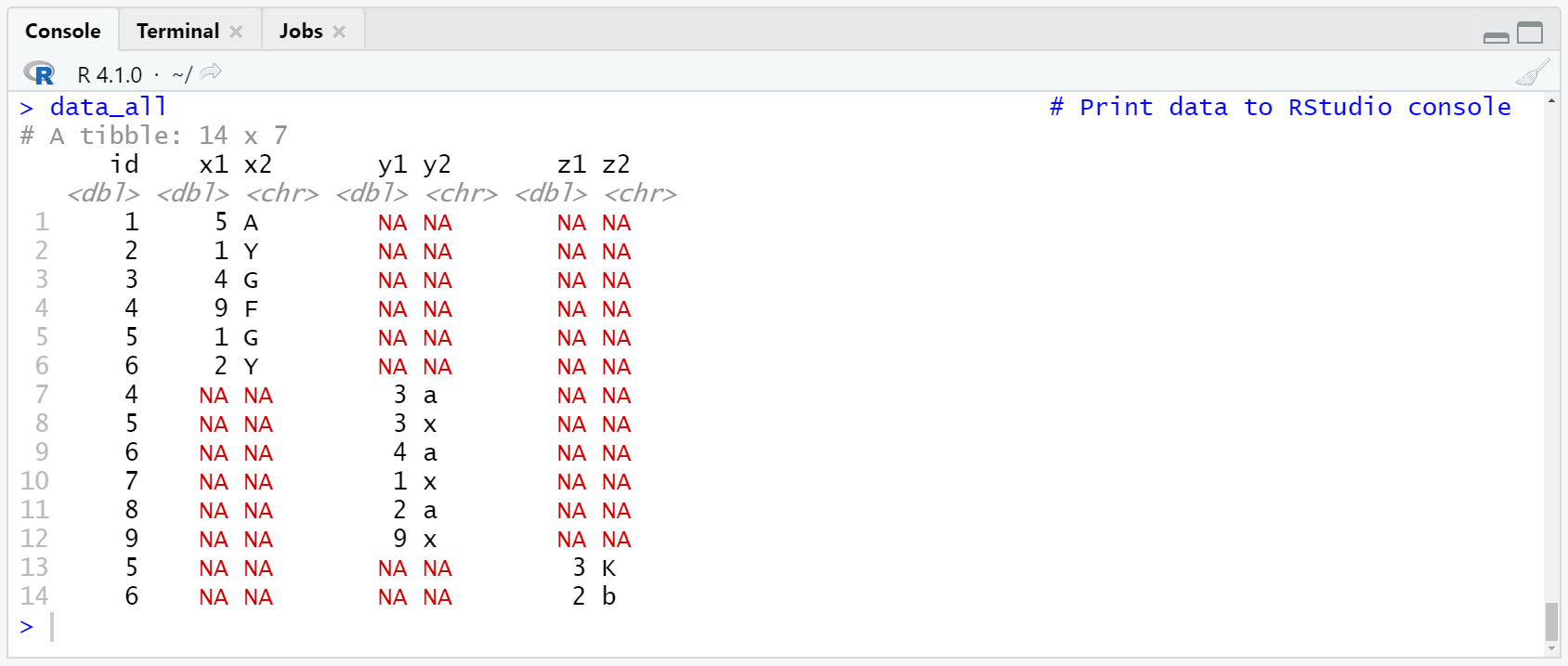
Table 1: Tibble Containing Three Data Sets.
Table 1 shows the output of the previous R code. As you can see, our three data sets were combined vertically in a single data set. Data set cells were set to NA, in case a variable was not included in all data sets.
Note that our previous R syntax created a tibble instead of a data frame. In case you prefer to work with data frames, you could simply convert this tibble to a data frame as follows:
as.data.frame(data_all) # Convert tibble to data.frame
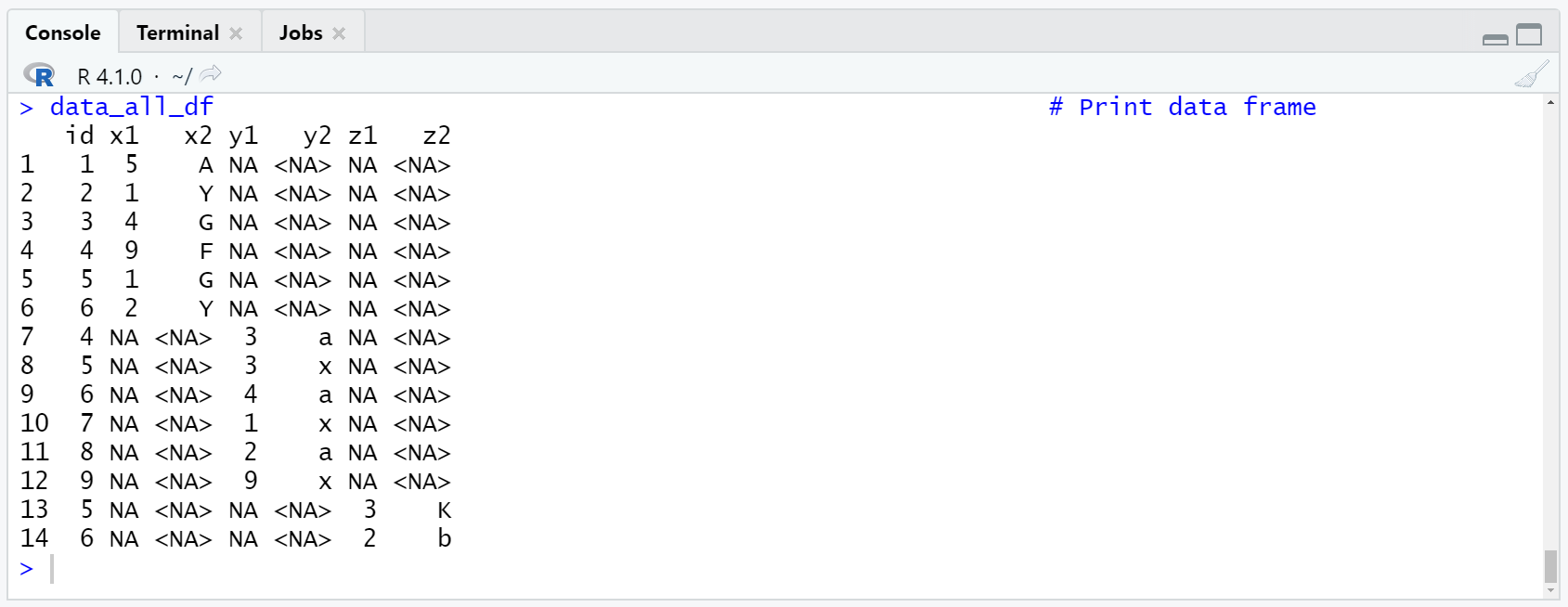
Table 2: Convert Tibble to Data Frame.
Looks good!
You may have noticed that we have simply stacked the rows of our three data frames on top of each other.
In the next example, I’ll explain how to merge our data frames based on a shared ID variable, so keep on reading!
Example 2: Import & Join CSV Files in R
As you can see in the previous tables, all of our example data frames contain an id column. In this example, we’ll use this id column to merge our data (instead of just row-binding it as in Example 1).
For this task, we first have to install and load the purrr package:
install.packages("purrr") # Install & load purrr package library("purrr")
Next, we can apply the reduce and full_join functions to join our data frames based in the id variables:
data_join <- list.files(path = "C:/Users/Joach/Desktop/my_folder", # Identify all CSV files pattern = "*.csv", full.names = TRUE) %>% lapply(read_csv) %>% # Store all files in list reduce(full_join, by = "id") # Full-join data sets into one data set data_join # Print data to RStudio console
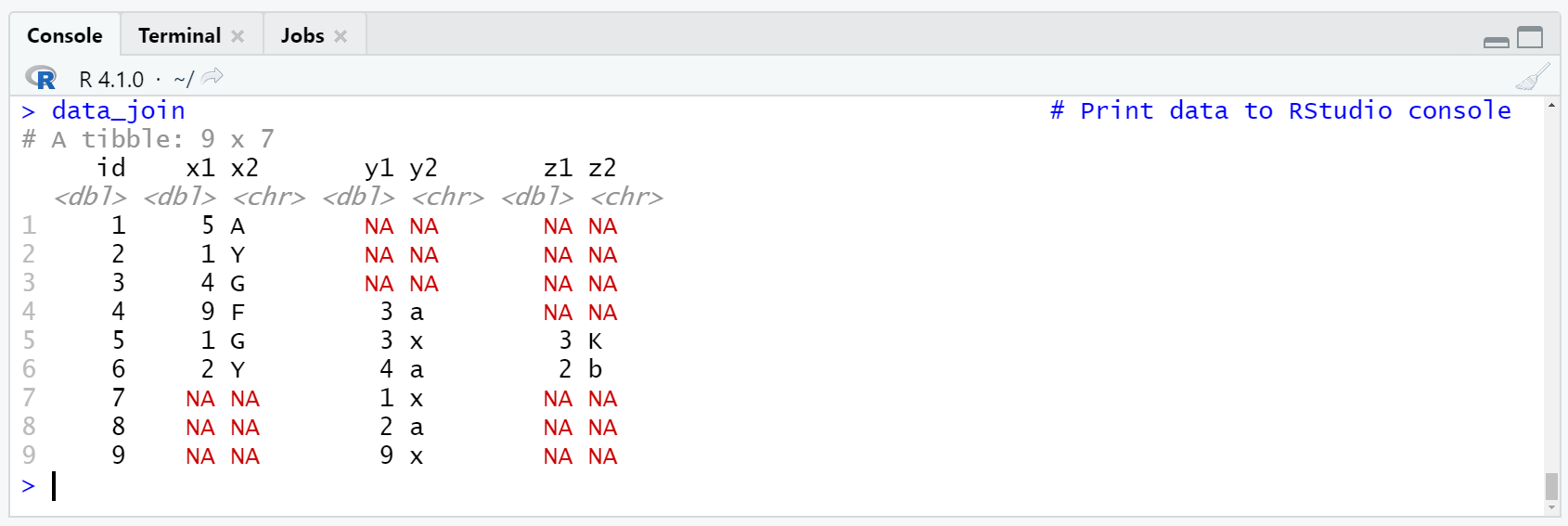
Table 3: Tibble Containing Three Data Sets Merged by ID Column.
As you can see based on Table 3, we have created another tibble containing our three data sets. However, in contrast to the previous example, we have joined our data sets based on the id column, and hence have avoided duplicated observations in our final data set.
Note that we have used a full join, to combine our data frames. In case you want to learn more about different types of joins, you may have a look here.
Video & Further Resources
Have a look at the following video that I have recently released on my Statistics Globe YouTube channel. I illustrate the contents of this article in more detail in the video.
In addition, I can recommend reading the other articles on this homepage.
- Read XLSX and XLS Files to R
- The dir Function in R
- Write & Read Multiple CSV Files Using for-Loop
- R Functions List (+ Examples)
- The R Programming Language
On this page I showed you how to combine all CSV files in a folder in the R programming language. Let me know in the comments section, in case you have further questions.
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.







30 Comments. Leave new
Hi Joachim,
First of all, thank you for the content.
I have tried to use this content to sort out one task, but no success.
Scenario:
I have a few csv files in a folder, I want to merge all of them using an apply function.
Step 1 – I tried first listing all the files from a folder.
Step 2 – Save all csv files from the step 1 to the work directory using an apply function. ( failed )
Step 3 – Merge all the files.
This is an easy process in Power BI using power query, I am trying to reproduce it using R.
Hey Fernando,
You are very welcome, thank you for the kind words!
Have you tried the example code of this tutorial? What kind of error message did you get? I need some more details 🙂
Regards,
Joachim
This is amazing, thank you so very much!
Thanks Lou, I’m glad to hear this!
Hello,
Thank you for this.
I tried your suggestion and what if I have one identical column for each file and I want all my files ordered by this identical column as with ‘by’ in merge (merge(data_1, data_2, by=c(“participant”, “X1”)))
Hi Alex,
You are very welcome!
Please have a look at the order function, specifically Example 5 of this tutorial: https://statisticsglobe.com/sort-order-rank-r-function-example
Regards,
Joachim
Hello Joachim,
Thank you for your answer but to merge 2 files with merge function, I can define a unique column as a reference for the merge (merge(data_1, data_2, by=c(“participant”, “X1”))). Here my 2 files will be merge according 2 specific and unique columns (with no duplication of my columns participant and X1). If I want to merge serveral csv with each a unique column that can give me the right association between all my file, is it possible ? I take a look on your example 5 (https://statisticsglobe.com/sort-order-rank-r-function-example) but it does not seem to answer my question. Until now, I am forced to use merge in an incremental way to merge all my csv because I specifically need the right association between my files with columns participant and X1.
Hey Alex,
Thank you for the follow up.
Does this (or something similar to this) work?
Regards,
Joachim
Yes, It worked. Thank you very very much!
Great to hear. You are very welcome Alex! 🙂
Joachim,
The csv files in my directory have the following problem:
for data1, variable x1, the class is numeric.
for data2, variable x1, the class is character.
example
data1 <- data.frame(id = 1:6, # Create first example data frame
x1 = c(5, 1, 4, 9, 1, 2),
x2 = c("A", "Y", "G", "F", "G", "Y"))
data2 <- data.frame(id = 4:9, # Create second example data frame
x1 = c("none", "none", "none", "none", "none", "none"),
y1 = c(3, 3, 4, 1, 2, 9),
y2 = c("a", "x", "a", "x", "a", "x"))
I want the new, combined dataset to have variable x1 look like this:
x1 = c(5, 1, 4, 9, 1, 2, NA, NA, NA, NA, NA, NA)
instead I get an error:
Error: Can't combine `x1` and `x1` .
Where/how in your series of pipe operators could I use the function as.numeric() on x1 variable – for all datasets in the list?
or na.strings = c(“none”)?
Thank you,
Tommy
Hi Tommy,
In this case I would do this:
1) Import the two data frames one-by-one
2) Harmonize data frame classes using the as.numeric function
3) Merge data frames
I hope that helps!
Joachim
How do it if I want to use sep = “\t” in my dataframe?
Hey Alberto,
I think you should be able to add additional arguments within the lapply function (i.e. lapply(read_csv, sep = “\t”).
Does this work for you?
Regards
Joachim
Hey..it was no doubt an awesome explanation. But what about the case where I want to read CSV files one by one and run few set of instructions on each file seperately and write the output seperately…something like loop through all files one by one …. Regards
Hey Amit,
Thank you for the kind feedback!
Indeed, a for-loop might be a good solution for you. The code could look like this:
You can find more info on for-loops here: https://statisticsglobe.com/loops-in-r/
Regards
Joachim
Dear Joachim,
First of all, thank you for the content. I have learnt a lof from this amazing blog and your videos. I am a Stata user but I am trying to move to R gradually.
This discussion about “Import & Merge Multiple csv Files” is really great. I have this particular situation. I have about 20 data files, one per year (2000-2019), with lots of health infotmation. The problem comes when those data change the number of columns, and the class of many variables whith time (numeric to character or viceversa for the same variable). The expected result is to have only one data frame for the whole period.
I’ve read your suggestions to some questions in this post and think that my main problem is to harmonize the data frame classes. Is there a way to declare the class of the variables in a temporal data frame (whitout data), and then start reading and merging each CSV file using a for-loop?
Thanks in advance for your feedback!!!
Keep doing this amazing work!!!
I’m your FAN!!!!
Regards,
Alberto
Hey Alberto,
Thank you very much for the very kind words! Glad to hear that you find my tutorials helpful!
Dealing with data classes when importing data from different sources and programming languages is always a mess. I have experienced this problem a lot myself (at my previous job were many STATA and SAS users).
Anyway, I have tried to replicate your situation and have written some code to import and rbind the data with proper classes:
I’m sure the efficiency of this code can be improved, but for the given example data it works fine.
I hope that helps! 🙂
Joachim
Hi ,
Thank for this useful blog, but i am facing some problems.
I have 200 csv files with each file having two columns, first is the plot ID which is same for all the 200 files, however the second column in each file consist of data with different variable name. I have run these codes but in results the data is merged in one column, however i need a data frame with data in columns for each variable.
Please help me in this regard.
Waiting for your reply.
Thank you
Hey,
Thank you for the kind words!
Could you share the code you have used?
Regards,
Joachim
Hi Joachim,
Your blog has helped so much with quickly getting up to pace in R! Thanks so much.
A quick questions though, I have roughly 20 csv files with the first column of each being an ID code (string). Is there a way to merge them in R so that the data for each ID code is kept assigned to that row?
Hey Sarah,
Thank you so much for the kind words, glad you find my tutorials helpful!
Also, thanks a lot for the question. I have noticed that this tutorial is a bit older/outdated, and therefore I have worked over it once again. You can now find a second example, which shows how to join the imported data sets based on their ID.
I hope this helps!
Joachim
Hello Joachim, my CSVs to be merged all have 12 header rows that can eventually be deleted, but first I have to create a new column for “SampleID”, then extract the SampleID from cell B2, and paste the SampleID value in all the rows from a given csv file, then the top 12 rows can be deleted. Would appreciate any suggestions.
Hey Joe,
Are you searching for something like this?
Regards,
Joachim
Hi there,
This little bit of code was super useful for me to combine a bunch of related csvs. Do you have a suggestion of where to add a bit of code to extract the file name and add it as a column to each file as they are imported? I need to be able to distinguish which data came from which file after I’ve combined them and they don’t have a unique identifier once the csvs are combined.
Hey Megan,
Thanks for the kind comment, it’s great to hear that this tutorial is helpful!
Regarding your question, please have a look at the example code below. I’m sure there must be a more efficient solution for this, but the following code should do what you want:
Regards,
Joachim
Thank-you, this worked great for me 🙂
This is great to hear, thanks for the kind comment Megan! 🙂
Hi Joachim,
I have 12 .csv files in a folder. I’m trying to combine all these files into one dataframe. They all have similar coumns. I tried using lapply and rbind methods. But when i try to view contents of a column it shows null.
Could you please tell me a method to solve that?
Hey Nikitha,
In order to combine data frames using rbind, the data sets need to have the exactly same column names. Is this the case?
If not, you may have a look at the rbind.fill function of the plyr package.
Regards,
Joachim