Calculate (Root) Mean Squared Error in R (5 Examples)
In this tutorial you’ll learn how to compute the mean squared error (MSE) and the root mean squared error (RMSE) in R programming.
The article consists of five examples for the computation of the MSE and RMSE. More precisely, the page consists of this information:
It’s time to dive into the examples!
Creating Example Data
As the first step, we have to construct some example data:
set.seed(39756934) # Create example data x <- rnorm(100) y <- rnorm(100) + 0.5 * x my_data <- data.frame(x, y) head(my_data) # Print head of example data
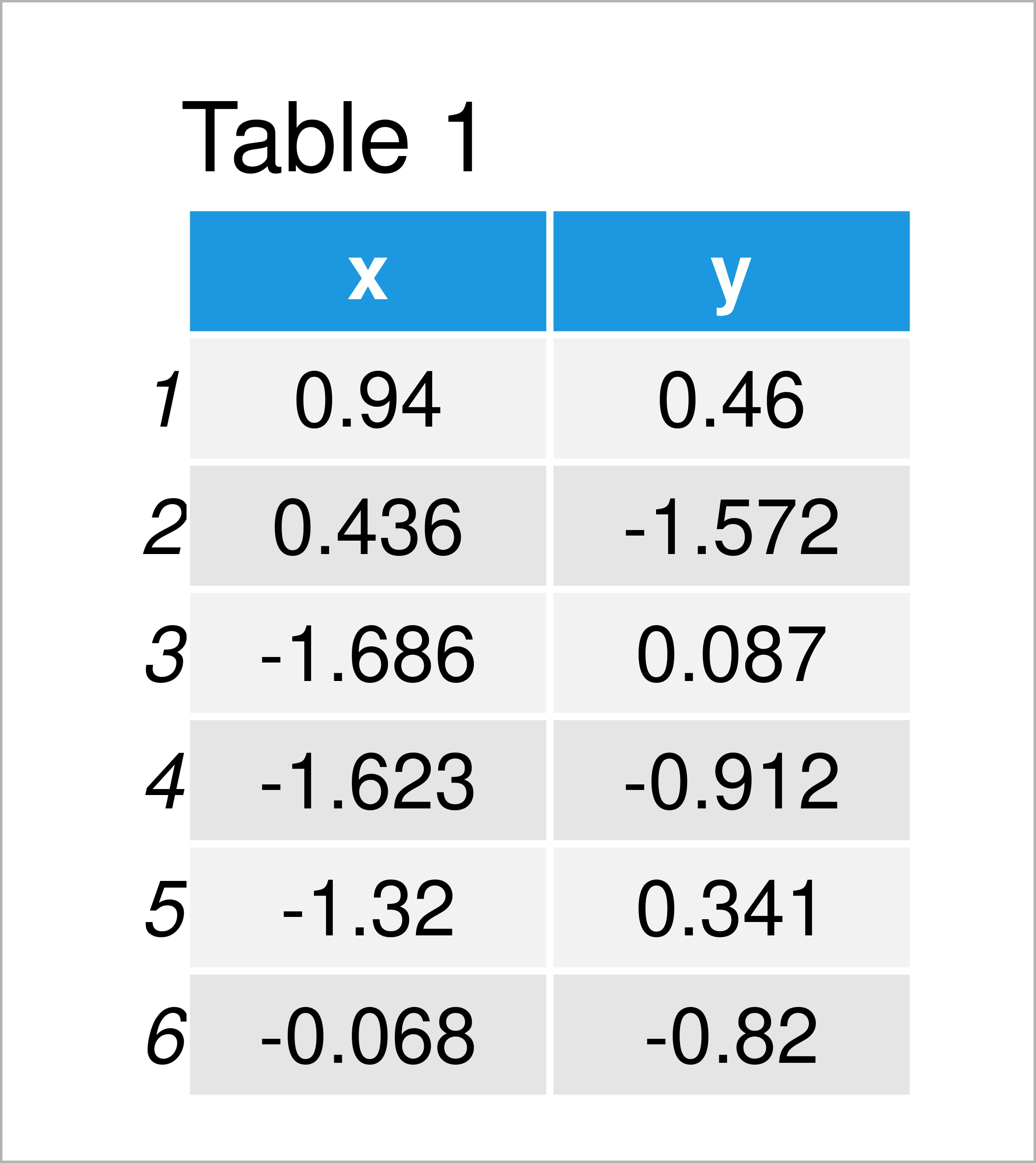
As you can see based on Table 1, our example data is a data frame consisting of the two columns “x” and “y”.
Next, we can estimate a linear regression model using the lm function:
my_mod <- lm(y ~ x, my_data) # Estimate linear model
The previous R code has created a new data object called my_mod, which contains the output of our linear regression. In the following examples, we’ll use this model object to compute the MSE and RMSE.
Example 1: Calculate MSE Using mean() Function & Residuals
Example 1 illustrates how to calculate the mean squared error based on the mean function and the residuals of our linear regression.
Have a look at the following R code and its output:
mean(my_mod$residuals^2) # Calculate MSE # [1] 0.7643822
As you can see based on the previous output of the RStudio console, the MSE of our analysis is 0.7643822.
Example 2: Calculate MSE Using mean() & predict() Functions
Example 1 has explained how to compute the MSE using the mean function and the residuals of our model. However, the R programming languages provides alternative ways to calculate the mean squared error.
This example illustrates how to use the mean and predict functions to calculate the MSE of a regression analysis:
mean((my_data$y - predict(my_mod))^2) # Calculate MSE # [1] 0.7643822
The result is exactly the same as in Example 1.
Example 3: Calculate MSE Using mse() Function of Metrics Package
So far, we have only used the functions provided by the basic installation of the R programming language.
Example 3 explains how to compute the MSE using the mse() function of the Metrics package.
First, we need to install and load the Metrics package.
install.packages("Metrics") # Install & load Metrics package library("Metrics")
Next, we can apply the mse and predict functions to calculate the MSE:
mse(my_data$y, predict(my_mod , my_data)) # Calculate MSE # [1] 0.7643822
Again, the result is 0.7643822. Whether you want to use the Metrics package or one of codes of the previous Examples is a matter of taste!
Example 4: Calculate RMSE Using mean() & sqrt() Functions
We can easily adjust the previous R codes to calculate the root mean squared error (RMSE) instead of the mean squared error (MSE).
For this task, we can simply apply the sqrt function to the output of one of the previous codes to calculate the square root of this result.
In this example, I’m applying the sqrt function to the R syntax of Example 1:
sqrt(mean(my_mod$residuals^2)) # Calculate RMSE [1] 0.8742895
As you can see, the RMSE of our regression model is 0.8742895.
Example 5: Calculate RMSE Using rmse() Function of Metrics Package
The Metrics package that we have already used in Example 3 also provides a function to calculate the RMSE. This function is called rmse() and can be applied as shown below:
rmse(my_data$y, predict(my_mod , my_data)) # Calculate RMSE [1] 0.8742895
The result is the same as in the previous example.
Video & Further Resources
I have recently published a video on my YouTube channel, which explains the examples of the present tutorial. You can find the video below.
In addition, you may want to have a look at the other articles on this website. Please find a selection of related articles below:
In summary: On this page you have learned how to calculate the (root) mean squared error in R programming. Tell me about it in the comments section, in case you have further questions and/or comments. Furthermore, please subscribe to my email newsletter in order to get updates on new articles.
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.







4 Comments. Leave new
I’m Adrian Keister, who left a comment on your YouTube – which appears to be malfunctioning and not letting me leave my comment. So here’s a MWE:
set.seed(39756934) # Create example data
x = rnorm(100)
y = rnorm(100) + 0.5 * x + 3
my_data = data.frame(x, y)
min(y)
0.6716385
my_mod = glm(y~x, data=my_data, family=Gamma())
mean(my_mod$residuals^2)
0.01388825
mean((my_data$y – predict(my_mod))^2)
6.954654
Hey Adrian,
I’ve just responded to your comment on YouTube. I’ll copy/paste my response here as well:
Thanks for sharing your code. Unfortunately, I don’t have experience in calculating the MSE for Gamma distributions, but I found this function which might be helpful for your case: https://www.rdocumentation.org/packages/DELTD/versions/2.6.5/topics/mseGamma
I hope this helps!
Joachim
Asked on SO.SE, and got the idea that both the residuals and the predict functions need to have type=”response” in order to ensure equality in more general cases.
Please see my response to your other comment.