R Error in lm.fit(x, y, offset, singular.ok , …) : 0 (non-NA) cases (2 Examples)
In this article, I’ll illustrate how to debug the “Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : 0 (non-NA) cases” in the R programming language.
The table of content is structured like this:
Let’s get started…
Example Data
Let’s first construct some example data:
set.seed(9364593) # Create example data data <- data.frame(y = rnorm(100), x1 = rnorm(100), x2 = NA) head(data) # Head of example data
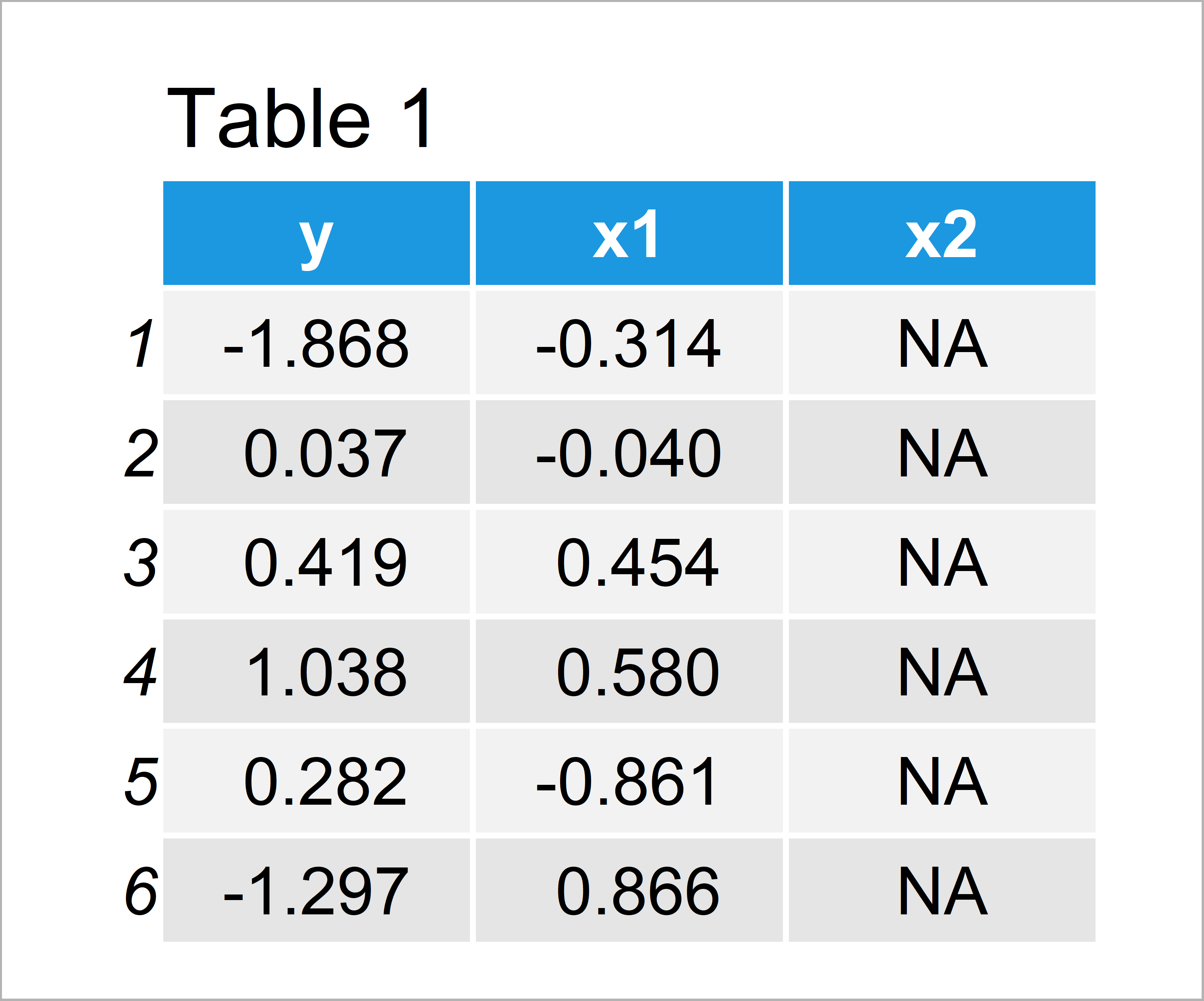
Have a look at the previous table. It reveals that our example data is constructed of 100 rows and three columns.
Example 1: Reproduce the Error in lm.fit – 0 (non-NA) cases
The following R syntax illustrates how to replicate the “Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : 0 (non-NA) cases” in the R programming language.
Let’s assume that we want to estimate a linear regression model using the lm function in R. Then, we might try to use the following R syntax:
lm(y ~ ., data) # Estimate model based on entire data set # Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, ...) : # 0 (non-NA) cases
Unfortunately, the RStudio console returns the error message “0 (non-NA) cases”.
The reason for this is that one (or multiple) of our data frame columns contains only NA values. The lm function (and other modelling functions as well) can not handle such only NA predictors.
So how can we solve this problem?
Example 2: Fix the Error in lm.fit – 0 (non-NA) cases
The following R code explains how to deal with the “Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : 0 (non-NA) cases”.
To avoid this message, we have to remove all independent variables that contain only missing values from our model.
In the following R code, I’m explicitly specifying that I want to use only the column x1 as predictor variable:
lm(y ~ x1, data) # Estimate model based on subset # Call: # lm(formula = y ~ x1, data = data) # # Coefficients: # (Intercept) x1 # -0.15534 0.06922
As you can see, the lm function has returned a valid output without any error messages.
Video, Further Resources & Summary
Do you want to learn more about errors? Then I recommend watching the following video of my YouTube channel. In the video, I’m explaining the R programming code of this article:
The YouTube video will be added soon.
Furthermore, you might want to read the other posts on this homepage.
- Help – Error in if (NA) { : missing value where TRUE/FALSE needed
- Dealing with Warnings & Errors in R (Example Codes)
- The R Programming Language
At this point you should know how to handle the “Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : 0 (non-NA) cases” in R. If you have any additional comments and/or questions, let me know in the comments section.







39 Comments. Leave new
how to handle when I still get this error despite having independent variables which have no NAs whatsoever?
Hi Rakshathi,
Could you illustrate how your data looks like and share your code?
Regards
Joachim
I want to make a linear reg with y, x1 and x2
with m1 <- lm(data=data1, y ~ x1, na.action = "na.exclude") I get a result
with m2 <- lm(data=data1, y ~ x2, na.action = "na.exclude") I get a result
with m3 <- lm(data=data1, y ~ x1, x2, na.action = "na.exclude") I get Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) :
0 (non-NA) cases
What happens here?
both x1 and x2 contain some NA, but only 20 of 140 observations
Hey,
It seems like the specification of your formula is incorrect. You would have to use a + sign between x1 and x2 in your third example:
Does this fix your error?
Regards,
Joachim
Hello!
I get the same error when trying to perform a two-ways anova.
My script:
res.aov <- anova_test(
data = Apportal, dv = score, wid = record_id,
within = c(time, visit))
Error message: Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) :
0 (non-NA) cases
Output: get_anova_table(res.aov)
ANOVA Table (type III tests)
Effect DFn DFd F p p<.05 ges
1 visit 1 29 1.68e-30 1 1.7e-33
I have NO missing values in any of my variables.
Whay am I doing wrong?
Thank you,
Bea
Hey Bea,
This is difficult to evaluate without seeing your data. Could you share (a subset of) the data here, or send it to joachim@statisticsglobe.com ?
Regards,
Joachim
Hi! I actually have the same problem as Bea, only with a Repeated Measures ANOVA. I have no missing values in my data
res.aov <- anova_test(
data = dat, dv = e.coli, wid = id,
within = c(site, time)
)
get_anova_table(res.aov)
Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) :
0 (non-NA) cases
and I don't even get an output…
id site time weight e.coli
1 p t1 0,2297 96968256,53
2 ww t1 0,2354 4575876,75
3 p t1 0,1959 35296621,32
I would appreciate any help.
Thanks! Lisa
Hey Lisa,
Apologies for the late reply, I just got back from vacation. Do you still need help with this question?
Regards,
Joachim
I’m having the exact same issue as Lisa, no NAs in my data frame but this error appears when I try to run a repeated measures ANOVA in R using the same method Lisa mentioned.
Hey Jess,
Unfortunately, I have no experience with this error message when using ANOVA myself. However, I found this thread on Stack Overflow, which seems to explain the problem.
I’m quoting from the answer of StupidWolf in this thread:
“For repeated measure anova, you need complete observations for each time point, before and after treatment”
Is this maybe the reason for your error?
Regards,
Joachim
Hi, Joachim
Hi everyone,
I did a diary study.
It was 5 days, 2 times a day (morning and afternoon).
Morning measures I have:
– sleeping problems
Afternoon measures I have:
– Emotions
– Incivility
head(base_within_merged,6)[,c(“code”,”register”,”sleepingproblems”,”expf2fwi”,”emotionalreactivity”)]
code register sleepingproblems expf2fwi emotionalreactivity
1 aaja28 7 1.8 NA NA
2 aaja28 8 NA 1 1.000000
3 aaja28 5 2.0 NA NA
4 aaja28 6 NA 1 2.666667
5 aaja28 10 NA NA NA
6 aaja28 3 2.6 NA NA
library (nlme)
library (lme4)
I want to test a mediation. First I want to test the model with just the IV. But, when I do this on R:
summary(m1 <- lmer(sleepingproblems ~ incivility + (1 + incivility |code), data = data_within_measures))
I get the error:
Error in h(simpleError(msg, call)) : **
** error in evaluating the argument 'object' in selecting a method for function 'summary': missing value where TRUE/FALSE needed
I get the error. The reason is that because in the morning I only have measures for SP and in the afternoon I only have data for incivility, so they don´t match. But I don´t know how to solve it.
I would like to know if there is a way I can match both (e.g. matching the data from day 1 morning with the afternoon data, and so on). The idea is to see whether who experiences incivility will most likely experience sleeping problems as well.
I appreciate the help!
Thank you!!
(I got a mistake in previous post)
Hi, Joachim
Hi everyone,
I did a diary study.
It was 5 days, 2 times a day (morning and afternoon).
Morning measures I have:
– sleeping problems
Afternoon measures I have:
– Emotions
– Incivility
head(base_within_merged,6)[,c(“code”,”register”,”sleepingproblems”,”expf2fwi”,”emotionalreactivity”)]
code register sleepingproblems expf2fwi emotionalreactivity
1 aaja28 7 1.8 NA NA
2 aaja28 8 NA 1 1.000000
3 aaja28 5 2.0 NA NA
4 aaja28 6 NA 1 2.666667
5 aaja28 10 NA NA NA
6 aaja28 3 2.6 NA NA
library (nlme)
library (lme4)
I want to test a mediation. First I want to test the model with just the IV. But, when I do this on R:
summary(m1 <- lmer(sleepingproblems ~ incivility + (1 + incivility |code), data = data_within_measures))
I get the error:
Error in lme4::lFormula(formula = sleepingproblems ~ expf2fwi + (1 | code), :
0 (non-NA) cases
I get the error. The reason is that because in the morning I only have measures for SP and in the afternoon I only have data for incivility, so they don´t match. But I don´t know how to solve it.
I would like to know if there is a way I can match both (e.g. matching the data from day 1 morning with the afternoon data, and so on). The idea is to see whether who experiences incivility will most likely experience sleeping problems as well.
I appreciate the help!
Thank you!!
Hello Francisca,
It looks like it is about how you merge your data in the beginning. Normally, if you merge by id properly, there shouldn’t be any missing values due to the reason you mentioned. But are you sure that NAs occur because of that? So this means you have duplicated ids or, in your case codes I guess. Maybe you have NAs because the data is indeed missing, then it’s another story. Also, what does register refer to?
Regards,
Cansu
Hi Cansu!
Thank you so much for trying helping me, really ☺️
Regarding the last question: registers are each point in time when they filled the questionnaires. So, they answered to 5 days, 2 times a day (a possible of 10 registers).
Registers 1,3,5,7,9 are for the questionnaires that were filled in the morning. 2,4,6,8,10 are for the registers that were filled in the afternoon.
So, imagine for sleeping problems I have values for registers 1,3,5,7,9 but I will not have it for 2,4,6,8 and 10 because these are the register for the afternoon questionnaires and I did not ask questions regarding sleeping problems in the afternoon.
Then, just regarding the merge:
I merged both databases from morning and afternoon in SPSS prior to start using RStudio.
The “data_within_measures” there is the merge I did of within dataset (morning +afternoon questionnaires) with the between-level dataset.
So, all in all, I want to test a mediation of Workplace Incivility ->emotions -> sleeping problems. And I first only wanted to go with a model of the IV(WI) and DV (SP), but it gives that error (in the previous comment).
Hello Francisca,
I think converting the data from the long to wide format would help. See my tutorial. The registers column should split into multiple columns per common id. Do you know which registers in the afternoon measure the emotions or incivility exactly? If so, my suggestion should help. If not, we might need some data manipulation that I should think about. Let me know about it.
Regards,
Cansu
Hi Cansu,
Thank you so much! I will try to do it (convert data from long to wide).
Regarding the registers, for sleeping problems there should be values for register 1,3,5,7 and 9.
For both emotions and incivility, as I asked about both in the afternoon I should have values for 2,4,6,8 and 10. Although, of course, not everyone answered to all the registers so there could be a case, that some people failed to answer to 1/2/3/4 registers, so there could be NA values in that case too.
So, the things is in the lines that I have values for sleeping problems (eg. register 1) I have NA values for emotions and incivility because there were any questions regarding these 2 variables in that particular questionnaire/moment.
But yes, anyway, I will follow your tutorial and put the data into the wide format and let you know how it works out.
Thank you so much, again.
Hello Francisca,
You are welcome. I asked if we know which registers are for the emotion measurements exactly, like 2, 4, and 6 measure the emotions, etc. That NAs occur due to non-response is a totally different story. So let’s see what you get first then, we can discuss how you can deal with the missing values.
Regards,
Cansu
Hi, Cansu, again!
So.. basically, 2 things:
1) I was checking, and you were absolutely right regarding the merging. It is not merging correctly, but the odd part is that merges well if I do it in one way, but not the other. I mean,
I coded:
base_within_merged <- merge(base_w_final,base_b[,c(18,120:135,137)],by="code") #base within with between-participant information
head(base_within_merged[,92:95],6)
sleepingproblems cognitivereapp exprsup emotionalregulation
1 1.8 3.500000 1 2.5
2 NA NA NA NA
3 2.0 3.500000 1 2.5
4 NA NA NA NA
5 NA NA NA NA
6 2.6 3.333333 2 2.8
But then I do:
attach(base_w_final)
aggdata <- aggregate(base_w_final[,c(1:3,92,95:101,105)], by=list(base_w_final$code), FUN=mean,na.rm=TRUE)
detach(base_w_final)
aggdata$code <- aggdata$Group.1
aggdata <- aggdata[,c(-1)]
base_b_merged <- merge(base_b[,c(18,120:127,131,132)],aggdata,by="code") #base between with within-participant information
head(base_b_merged[,14:22],6)
sleepingproblems emotionalregulation negaffect posaffect expf2fwi perpf2fwi expcyberwi
1 2.133333 2.600 1.866667 2.233333 1.083333 1.416667 1.00
2 1.550000 3.075 1.000000 1.650000 2.600000 1.850000 1.65
3 1.400000 3.740 1.020000 1.820000 2.050000 1.750000 2.00
4 1.400000 3.580 1.460000 2.700000 1.850000 1.350000 1.30
5 1.440000 3.580 1.020000 1.240000 2.000000 2.000000 2.00
6 1.240000 3.640 1.240000 2.200000 1.150000 1.000000 1.00
perpcyberwi emotionalreactivity
1 1.00 2.444444
2 1.05 2.000000
3 2.00 1.622222
4 1.35 2.488889
5 2.00 2.044444
6 1.00 1.133333
In the 2nd, the NAs disappear. I am not sure why it works for 1 and not the other. I am not sure also if it is ok to use the "base_b_merged".
Despite all this, I tried to go further and for example predict the intercept variation of Incivility on Sleeping Problems, but it does not work.
model.5 <- lme(sleepingproblems~register+expf2fwi, random=~1+register|code,
data=base_b_final,
control=list(opt="optim"))
Error in MEestimate(lmeSt, grps) :
Singularity in backsolve at level 0, block 1
In addition: There were 50 or more warnings (use warnings() to see the first 50)
So, I did not move further. I am not sure if the merging before is not alllowing me to go further.
I tried as well to reshape as we talked about, but I am not sure it does something as well. I put:
dailywide <- reshape(data=base_w_final,
timevar=c("register"),
idvar="code",
v.names= c("sleepingproblems","emotionalregulation","negaffect","posaffect",
"expf2fwi","perpf2fwi","expcyberwi","perpcyberwi","emotionalreactivity"),
direction="wide")
I got:
A tibble: 6 × 9
`sleepingproblems.c(1, 2, …` `emotionalregu…` `negaffect.c(1…` `posaffect.c(1…` `expf2fwi.c(1,…`
1 NA NA NA NA NA
2 NA NA NA NA NA
3 NA NA NA NA NA
4 NA NA NA NA NA
5 NA NA NA NA NA
6 NA NA NA NA NA
# … with 4 more variables: `perpf2fwi.c(1, 2, 3, 4, 5, 6, 7, 8, 9, 10)` ,
# `expcyberwi.c(1, 2, 3, 4, 5, 6, 7, 8, 9, 10)` ,
# `perpcyberwi.c(1, 2, 3, 4, 5, 6, 7, 8, 9, 10)` ,
# `emotionalreactivity.c(1, 2, 3, 4, 5, 6, 7, 8, 9, 10)`
>
All in all, I am really sorry for taking so much of your time. I am I am honestly doing the best I can, but as I have been doing this without any support and it has been a battle.
Hello Franncisca,
I think what you do in the second merge is taking the mean by code, so you do not only merge, you also summarise statistics. Correct me if I am wrong. That’s why NAs are omitted during averaging. It may look like you are far from the solution, but no worries, I guess it is all about getting the data in a proper shape, which is always the hardest part of statistical analysis. So I suggest you share your initial datasets (before merging) with me. Please try showing at least 10-15 rows per dataset; then, I can help you to merge properly. With the limited info at hand, I can’t help you further.
Regards,
Cansu
Hi Cansu,
Of course. I just used the “important” variables here, so it doesn´t become too confusing.
The model is basically to see:
Experienced incivility -> Emotional Reactivity -> Sleeping Problems
Sleeping Problems -> Emotional Regulation -> Perpetrated Incivility
For the base_b (between data) I put here my single measure of experienced face-to-face and online incivility (ge_wi_f_1/ge_wi_cy_1), my single measure for perpetrated face-to-face and online incivility (ge_wi_f_2/ge_wi_cy_2), sleeping problems (SP), my 2 variables for emotions and the codes that each person created when answering to the questionnaire.
data.frame(
ge_wi_f_1 = c(“2″,”3″,”2″,”1″,”2”,
“3”,”4″,”2″,”2″,”2″,”4″,”3″,”1″,”3″,”4″),
ge_wi_f_2 = c(“1″,”1″,”1″,”1″,”1”,
“1”,”1″,”2″,”1″,”2″,”2″,”2″,”1″,”2″,”2″),
ge_wi_cy_1 = c(“2″,”1″,”1″,”1″,”1”,
“1”,”3″,”1″,”1″,”2″,”1″,”3″,”1″,”3″,”1″),
ge_wi_cy_2 = c(“1″,”1″,”1″,”1″,”1”,
“1”,”1″,”1″,”1″,”2″,”1″,”2″,”1″,”2″,”1″),
SP = c(1.2,1.6,1.6,2.4,2,
1.4,1,1.6,1.8,1.4,2,2.4,1.6,2,2.2),
EREA = c(2.77777777777778,
1.88888888888889,1.88888888888889,2.22222222222222,
2.22222222222222,2.22222222222222,2.22222222222222,
3.33333333333333,1.55555555555556,2.33333333333333,
3.44444444444444,3.22222222222222,2.55555555555556,
3.44444444444444,2.11111111111111),
EMOTIONALREGULATION = c(3.1,3.7,3.4,3.4,3.5,
3.5,2.7,2.6,3.2,3.2,3.1,3.4,3.8,3.2,4.2),
code = as.factor(c(“aaja28″,”abab29″,”abju01″,”acfe22″,”alju12”,
“amou79″,”amou91″,”baab81″,”baju47”,
“bano21″,”base85″,”beju58″,”bose99”,
“brno70″,”cade61”))
)
Regarding my base for within measures:
data.frame(
expf2fwi = c(NA,NA,2.5,NA,NA,1.25,
NA,1,NA,1,1.5,NA,3,NA,3.75),
perpf2fwi = c(NA,NA,2.5,NA,NA,1.25,
NA,2,NA,1,1.5,NA,1.25,NA,1.5),
expcyberwi = c(NA,NA,1.5,NA,NA,1,
NA,1,NA,1,1,NA,1,NA,1.75),
perpcyberwi = c(NA,NA,3,NA,NA,1,NA,
1,NA,1,1.25,NA,1,NA,1),
sleepingproblems = c(1.2,1.2,NA,2.2,2.6,
NA,2,NA,1.8,NA,NA,1.2,NA,2.2,NA),
emotionalreactivity = c(NA,NA,3.11111111111111,
NA,NA,3.66666666666667,NA,2.66666666666667,NA,1,
1.44444444444444,NA,1.44444444444444,NA,
3.11111111111111),
emotionalregulation = c(2.6,3.1,NA,2.9,2.8,
NA,2.5,NA,2.5,NA,NA,2.6,NA,3.6,NA),
code = as.factor(c(“152205074″,”152205074″,”152205074”,
“152205074”,”aaja28″,”aaja28″,”aaja28″,”aaja28″,
“aaja28″,”aaja28″,”abab29″,”abab29”,
“abab29″,”abab29″,”abab29”))
)
I think this is what you need, but if you need anything further please say so.
Thank you so much, again
Hello Francisca,
Thank you for sharing the datasets. So I need some info regarding the data collection and the experiment design. I understand that the measurements were taken for each individual once in the first dataset; is that correct? What is the aim of collecting this data? The second dataset has repeated measurements for each individual, I see. The same question arises here, what is the goal? I am confused that both datasets have some common variables: sleeping problems, emotional regulation, and code. It makes sense that both sets have the code variable represent the individuals. Still, I can’t get the difference between the variables of sleeping problems, emotional regulation, and emotional reactivity across the different datasets. Maybe you explain the experiment design a bit regarding this. And what happened to the registration variable, I can’t see it.
Regards,
Cansu
Hi Cansu,
Of course. You are absolutely right. I should explain better.
Basically, I did a diary study for 5 days, 2 times a day- one questionnaire in the morning, and one questionnaire in the afternoon.
Prior to the daily questionnaires, one week before, I did a general questionnaire, which is what I call my between-level data. In this general questionnaire I had the same variables as in the daily questionnaires, plus some others like sociodemographics, and other variables that I don´t expect much variability (e.g. questions regarding organizational culture). This general questionnaire was bigger, but answered only once, and I asked people If they had experienced any of what I was asking 3 months prior to that questionnaire- e.g. for incivility I asked, “in the past 3 months have you been in a situation where people ignored you…”.
Because they only answered once, I don´t have a register column for my base between-level. As well, in this general questionnaire, I asked people to create a code that would later be used for the daily questionnaires.
Concerning the daily questionnaires, the morning questionnaires, had to be answered any time between 8am and 10am. The afternoon questionnaires had to be answered any time between 4pm and 8pm.
In the morning I asked about the sleeping problems the night before, and about emotional regulation they were experiencing (at the moment they were answering). In the afternoon, I did not ask about sleeping problems because it wouldn´t make sense, nor I asked about emotional regulation again. Instead, I asked about if they experienced or perpetrated incivility throughout that specific day, I asked about emotional reactivity and others, which for the model are not important now.
What I want to test is: Experienced incivility -> Emotional Reactivity -> Sleeping Problems
Sleeping Problems -> Emotional Regulation -> Perpetrated Incivility
And basically, I want to see if on the one hand, if a person experiences incivility, will consequently be more emotional reactive and experience sleeping problems on that night. And, on the other hand, to see if one experiences sleeping problems the night before will lead to a lack of emotional regulation that in turn will lead to the perpetuation of uncivil behaviours towards others on the next day.
Please tell me if I did not explain myself well, of if there is any important information missing.
Thank you again for all the help 🙂
Hello Francisca,
Thank you for the information. First, I want to let you know that I have not evaluated your model or your research question. If you get any errors after merging the data while running the model, then we can discuss them. So, for now, I will assume the data is properly collected for such a model. I have two more questions before helping you with merging the datasets. Where is the registration info in the second dataset? I couldn’t see it. And you want to merge these sets for the matching codes, or in other words, individuals, is that correct?
Regards,
Cansu
Dear Cansu,
Of course. Thank you!
I leave here the data with the registers. Regarding the second questions, yes. The idea is to merge for the matching codes (each individual).
Here it is:
data.frame(
register = c(5, 7, 8, 9, 3, 4, 5, 6, 7, 8, 2, 3, 4, 5, 6),
expf2fwi = c(NA,NA,2.5,NA,NA,1.25,
NA,1,NA,1,1.5,NA,3,NA,3.75),
perpf2fwi = c(NA,NA,2.5,NA,NA,1.25,
NA,2,NA,1,1.5,NA,1.25,NA,1.5),
expcyberwi = c(NA,NA,1.5,NA,NA,1,
NA,1,NA,1,1,NA,1,NA,1.75),
perpcyberwi = c(NA,NA,3,NA,NA,1,NA,
1,NA,1,1.25,NA,1,NA,1),
sleepingproblems = c(1.2,1.2,NA,2.2,2.6,
NA,2,NA,1.8,NA,NA,1.2,NA,2.2,NA),
emotionalreactivity = c(NA,NA,3.11111111111111,
NA,NA,3.66666666666667,NA,2.66666666666667,NA,1,
1.44444444444444,NA,1.44444444444444,NA,
3.11111111111111),
emotionalregulation = c(2.6,3.1,NA,2.9,2.8,
NA,2.5,NA,2.5,NA,NA,2.6,NA,3.6,NA),
code = as.factor(c(“152205074″,”152205074″,”152205074”,
“152205074”,”aaja28″,”aaja28″,”aaja28″,”aaja28″,
“aaja28″,”aaja28″,”abab29″,”abab29”,
“abab29″,”abab29″,”abab29”))
)
Thank you!
Hello Francisca,
I named your first data as data1 and the second data as data2. Then to avoid confusion, I subset only the matching codes. Because, as you provided only a snapshot, there were missing codes. I think the proper way of merging these datasets is as follows.
Maybe it is the same as what you have done; then it means you should check the requirements for the data structure to run a mixed model via lme(). Maybe you should change the structure of your data as we discussed earlier: wide/long. Like spreading the repeated variables over the columns, etc. I suggest you search for the error you get or check the sample datasets used for running this type of mixed model with multiple between and within effects. Btw I changed the variable names a bit to make it clear for myself, I hope you can track it.
Regards,
Cansu
Dear Cansu,
Yes.. I did not work.
And if you see from the merge you show here, there are still NA values (when merging the base between and within).
I am trying to reshape the database from long to wide, and I get this:
data.frame(
stringsAsFactors = FALSE,
NA,
row.names = c(“1”,
“11”,”21″,”31″,
“41”,”51″,”61″,
“71”,”81″,”90″,”100″,
“110”,”120″,
“130”,”140″),
day = c(1,1,
1,1,1,1,1,1,
1,1,1,1,1,1,1),
sleepingproblems.1 = c(NA,
NA,NA,1.8,1.4,
1.2,1.2,1.2,NA,1.6,
1.8,1.4,2.4,1.4,
NA),
emotionalregulation.1 = c(NA,
NA,NA,3.9,3.6,
3.4,3.8,2.3,NA,3.4,
3.2,3.1,3.2,3.9,
NA),
negaffect.1 = c(NA,
NA,NA,1.1,1.9,1,
1.7,1.2,NA,1.7,
1.2,1.2,1.3,1.9,
NA),
posaffect.1 = c(NA,
NA,NA,2.5,3,1.6,
3,2.5,NA,2.7,
1.4,2.8,1.5,3.4,NA),
expf2fwi.1 = c(NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA),
perpf2fwi.1 = c(NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA),
expcyberwi.1 = c(NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA),
perpcyberwi.1 = c(NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA),
emotionalreactivity.1 = c(NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA),
sleepingproblems.2 = c(NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA),
emotionalregulation.2 = c(NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA),
negaffect.2 = c(NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA),
posaffect.2 = c(NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA,NA,
NA,NA,NA,NA),
expf2fwi.2 = c(NA,
NA,1.5,2.25,2.25,
2,1,1.25,1,1,
2.5,2.25,2.75,1.5,
1.75),
code = as.factor(c(“152205074”,
“aaja28″,”abab29”,
“abju01″,”acfe22”,
“alju12″,”amou79”,
“amou91″,”baab81”,
“baju47″,”bano21”,
“base85″,”beju58”,
“bose99″,”brno70”))
))
So, basically, now I have for each code each variable for each registration- eg. sleepingproblems.1, sleepingproblems.2, (…) sleeping.problems.10. But now what do I do with it? Do I have to make a mean now for each variable for the 10 registrations (i.e. should I make a new column for sleeping problems, incivility, emotional reactivity where I make a mean across the 10 registrations?)
Because I would like to test the model and say:
summary(m1 <- lmer(sleepingproblems.1,sleepingproblems.3,sleepingproblems.5,sleepingproblems.7,sleepingproblems.9 ~ incivility.2,incivility.4,incivility.6,incivility.8,incivility.10 + (1 + incivility.2,incivility.4,incivility.6,incivility.8,incivility.10|code), data = data_within_merged))
Thank you! 🙂
Hello Francisca,
I have never worked with mixed models in R with such high dimensional data. Therefore, I am afraid that I can’t help you with the model. Do you still get the same error while using the wide dataset? If so, as I suggested earlier, you should search for the error you get and find the reason why you got it, and fix it accordingly. You can not expect that the missing values will disappear, it makes sense that they still exist, but maybe you should check what missingness handling methods are offered by the lmer method or you can apply multiple imputation if you assume that the missingness is MAR. You should also check if your data is in the right format to run a mixed model; you should check the examples on the internet. For instance, I found this datacamp video by a quick search. Alternatively, you can run a simpler model to see if you can get a proper output and then adapt it to your original model. Let’s work on your model more, obtain a no-warning result, and find me here to let me know about your progress! If you still feel stuck, post your case on our Facebook discussion group. Ps: Taking average is not a good idea; you should keep your raw data.
Regards,
Cansu
Hi Cansu,
Thank you for your answer!
I can run the model if the variables are on the same point in time- e.g. sleeping problems with emotional regulation. But I always get an error if they are not (due to the NA).
I seem to have done the reshape of data to wide format well, but my problem is that I don´t know how to test the lme/lmer function in this format, and I don´t seem to find an explanation anywhere (I only seem people saying to use the long format). Part of my problem in testing in the wider format is that I used like this:
model1 <- lmer(sleepingproblems.1+sleepingproblems.3+sleepingproblems.5+sleepingproblems.7+sleepingproblems.9 ~
expf2fwi.2+ expf2fwi.4+ expf2fwi.6+ expf2fwi.8+ expf2fwi.10+ (1 + expf2fwi.2+ expf2fwi.4+ expf2fwi.6+ expf2fwi.8+ expf2fwi.10|code), data = merged_wide)
But then, I get an error saying:
Error: number of levels of each grouping factor must be < number of observations (problems: code)
Which I understand because I cannot group the code because my data is in the wide format.
Anyway… I will see and try my best.
Thank you!
Francisca
Hello Francisca,
Okay, if people say you should use the data in a long format, then keep it long. Also, if you are sure that your problem is due to NAs, then search what kind of dealing methods that lmer offers. Usually, there is a default method (even if it is not the best one) that’s why I was surprised that it gives an error. For instance, in the documentation, it says that listwise deletion (na.omit()) is the default. You can check the other options for the na.action argument, see the documentation. And as I suggested before, try to simplify your model and run it without any errors. Or you can even impute the missing values arbitrarily, let’s say, give 1s to all NAs. (This is not a proper way of imputing data or so, just for testing). Then run the model and see if it gives a solution or if it still gives an error. This is also a way of testing if you set your model wisely. Because it still feels me like the issue is due to the model, not the Nas directly. But I make a guess based on limited info and knowledge. As an option, you can always post your case on our Facebook discussion group: https://www.facebook.com/groups/statisticsglobe.
Regards,
Cansu
Cansu , I am writing this in another comment, but if there is in any way I can thank you for your support (monetarily, for instance), please say so.
It can be like a tutoring, or some sort of help.
I just think with this support it could make sense.
Francisca
Hello Francisca,
I am flattered, but we do not provide any private support/tutoring. Answering comments is one of my daily tasks, so no worries. I will try to help with merging when I have time; then the rest is your job 🙂 I just wanted to do my best, as it is a fundamental step to carry on the analysis, and it is a matching topic with our tutorials on Statistics Globe.
Regards,
Cansu
Thank you so much 🙂
I think is so hard to find support on these matters, that when there is it is like it worthy of some sort of “pay”.
Anyway, if you change your mind, feel free to say.
Regarding the merging, of course 🙂 If there is anything I can do or provide please say so.
Thank you again,
Francisca
You are welcome, if I need something further regarding merging, I will contact you on the other comment thread. Have a good one!
Regards,
Cansu
Hi Cansu,
I replaced all values with 1 just for the sake of testing and it works. I tried testing with the NAs and use “na.action” but still doesn´t work.
I am not sure if this say anything.
Thank you!
Francisca
Hello Francisca,
Then you should be sure that if your model and the experiment design are compatible. Maybe as I suggested, you simplify your model and try again. That should help to understand the problem. Also please describe your model for one last time, and then I will take a last look at it considering the dataset. If I can come up with something, then we discuss it; if not, you can consult our Facebook discussion group. Also, you can forget about handling the missingness methods because the missingness is due to the design, as you said before.
Regards,
Cansu
Regards,
Cansu
Hi Cansu,
I was trying to do Multiple Imputation as you suggested earlier, but I am not getting success.
Anyway, regarding the model:
What I want to test is: Experienced incivility -> Emotional Reactivity -> Sleeping Problems
Sleeping Problems -> Emotional Regulation -> Perpetrated Incivility
And basically, I want to see if on the one hand, if a person experiences incivility, will consequently be more emotional reactive and experience sleeping problems on that night. And, on the other hand, to see if one experiences sleeping problems the night before will lead to a lack of emotional regulation that in turn will lead to the perpetuation of uncivil behaviours towards others on the next day.
Morning measures (register 1,3,5,7,9): Sleeping problems, emotional regulation
Afternoon measures (register 2,4,6,8,10): incivility, emotional reacitivity
Thank you for the final look!
I know this is a nightmare.
Francisca
Hello Francisca,
Yes, the imputation suggestion was not a good one. As I dealt with many comments simultaneously, I lost the context a bit. Also, I couldn’t come up with any extra thoughts base don your recent explanation. I have never worked on testing such effects. I think you need help from someone more experienced. I think you explain the model well above. If you also share your data with the variable explanations on our Facebook discussion page, someone might help. Also, consider other discussion platforms like StackOverflow, etc. Sorry for not being helpful enough at this.
Regards,
Cansu
Hi Cansu!
I really appreciate your comments, and help regardless of the outcomes.
The only reason I don´t put it in Facebook is because I did it recently in a big RStudio group and I got lots of different people with fake profiles trying to extort me money and all that.
I will try to see if I get help from someone! 🙂
You are welcome. I wish you the best of luck!
Regards,
Cansu