Create Heatmap in R (3 Examples) | Base R, ggplot2 & plotly Package
In this post you’ll learn how to draw heatmaps in the R programming language.
The content of the article is structured as follows:
- Construction of Example Data
- Example 1: Create Heatmap with heatmap Function [Base R]
- Example 2: Create Heatmap with geom_tile Function [ggplot2 Package]
- Example 3: Create Heatmap with plot_ly Function [plotly Package]
- Video & Further Resources
Let’s dive right in.
Construction of Example Data
We’ll use the following matrix for the examples of this R tutorial:
set.seed(123) # Set seed for reproducibility data <- matrix(rnorm(100, 0, 10), nrow = 10, ncol = 10) # Create example data colnames(data) <- paste0("col", 1:10) # Column names rownames(data) <- paste0("row", 1:10) # Row names
Our data contains ten columns and ten rows with normally distributed random values.
In the following examples, I’ll show how to create heatmaps in R based on different functions and packages. It’s useful to know different ways to create heatmaps, since every package provides a different heatmap design. So keep on reading until the end of the article!
Example 1: Create Heatmap with heatmap Function [Base R]
The most common function for creating heatmaps in R is the heatmap() function, which is already provided by the base installation of R.
The heatmap function is applied as shown below:
heatmap(data) # Apply heatmap function

Figure 1: Default Heatmap in Base R.
Figure 1 illustrates the output of the previous R code. By default, the heatmap function returns a heatmap with column and row names as well as a dendrogram.
If we want, we can disable the automatically created dendrogram:
heatmap(data, Rowv = NA, Colv = NA) # Remove dendrogram
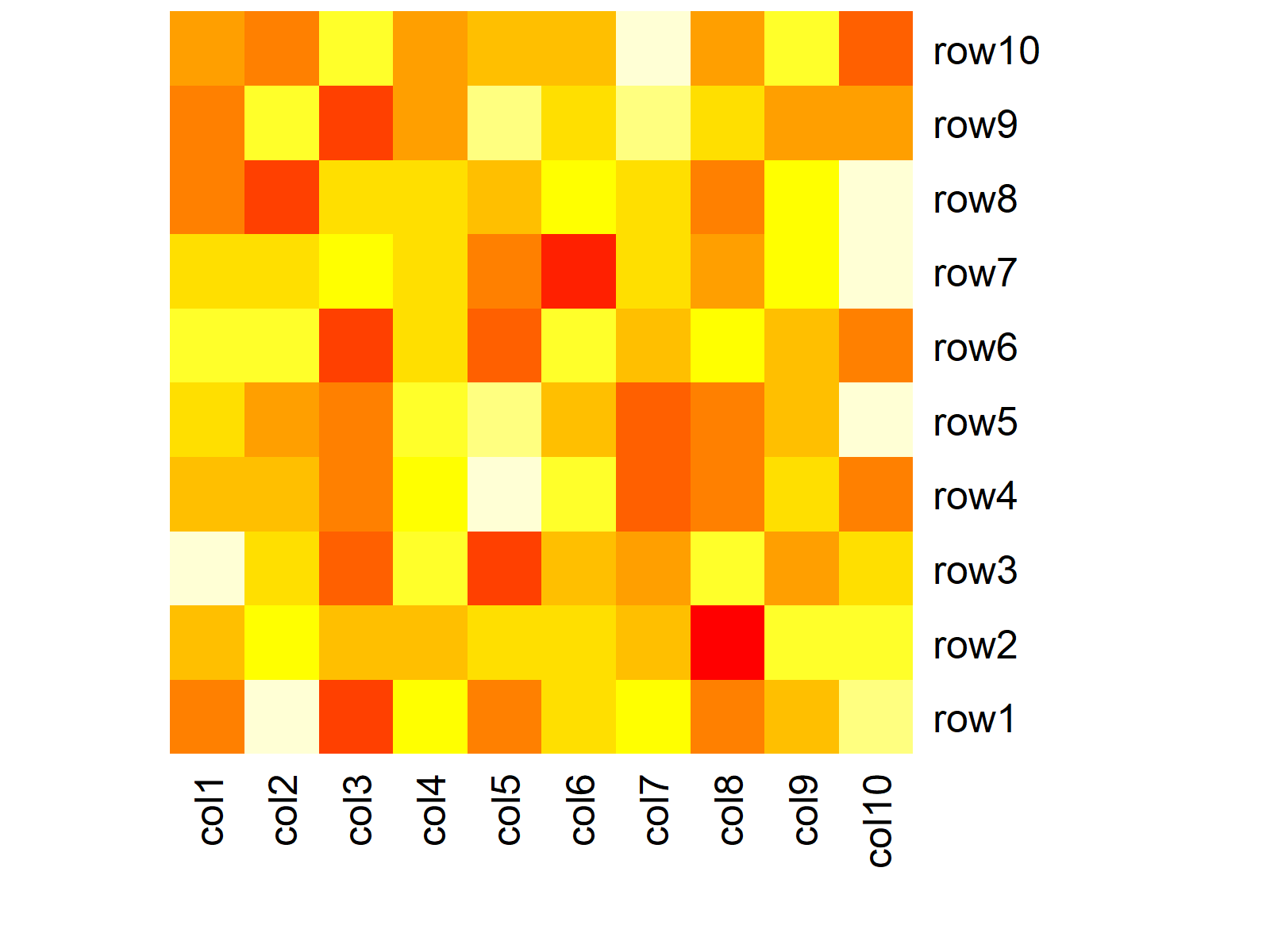
Figure 2: Heatmap without Dendrogram in Base R.
Furthermore, we can modify the colors of the heatmap by specifying our own color range with the colorRampPalette function. The following R code produces a function, which creates color ranges between the colors cyan and deeppink3:
my_colors <- colorRampPalette(c("cyan", "deeppink3")) # Manual color range
Now, we can create a heatmap with this color range as follows:
heatmap(data, col = my_colors(100)) # Heatmap with manual colors
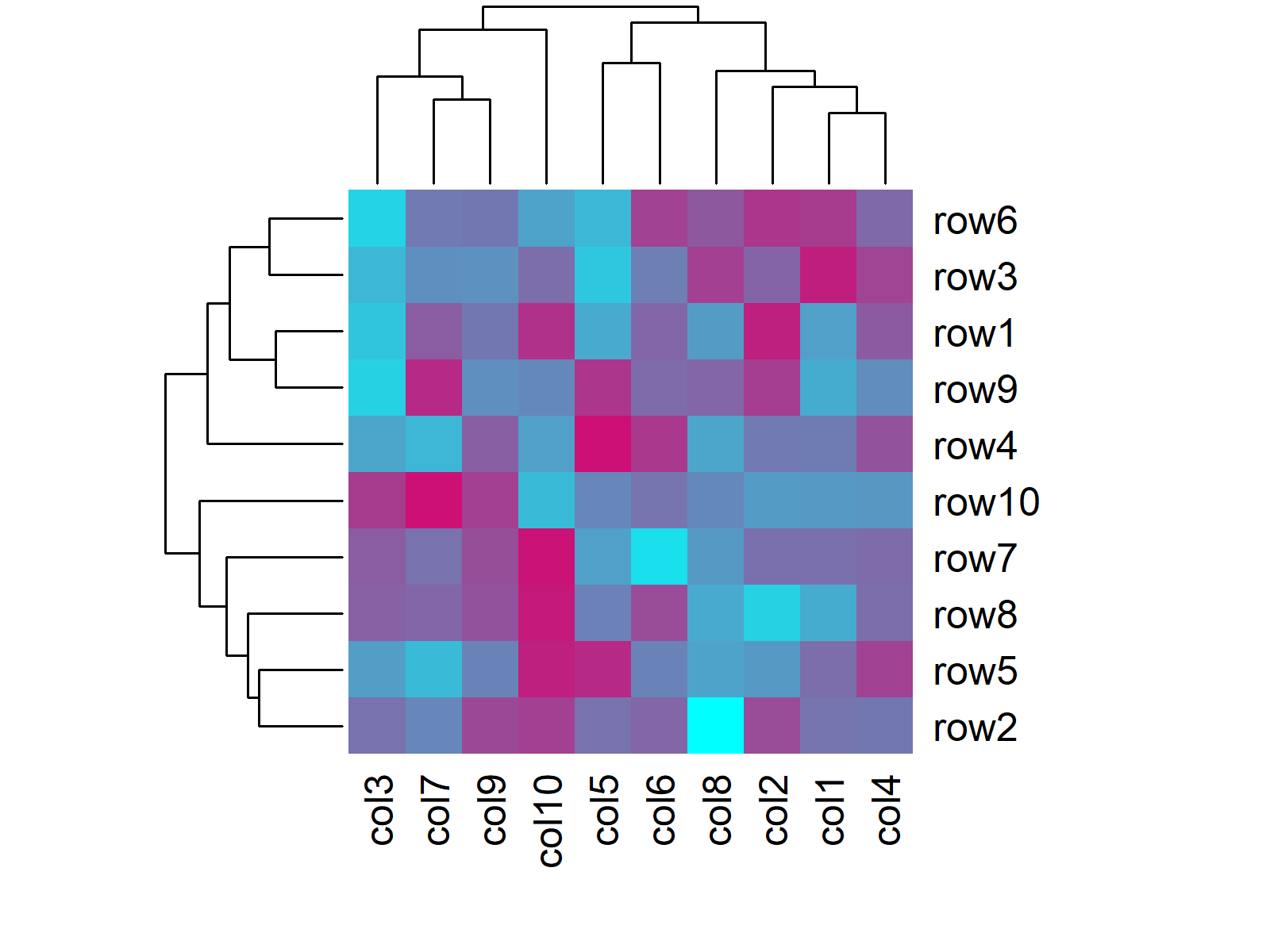
Figure 3: Heatmap with Manual Color Range in Base R.
Example 2: Create Heatmap with geom_tile Function [ggplot2 Package]
As already mentioned in the beginning of this page, many R packages are providing functions for the creation of heatmaps in R.
A popular package for graphics is the ggplot2 package of the tidyverse and in this example I’ll show you how to create a heatmap with ggplot2.
The ggplot2 package requires a long data format. We can create this data format with the reshape package…
install.packages("reshape") # Install reshape package library("reshape") # Load reshape package
…and the melt function:
data_melt <- melt(data) # Reorder data head(data_melt) # First six rows of data # X1 X2 value # row1 col1 -14.15254153 # row2 col1 -6.88199228 # row3 col1 2.44972363 # row4 col1 -0.04076637 # row5 col1 3.54613210 # row6 col1 -2.57462492
The RStudio console output shows the first 6 rows of our reshaped data. As you can see, the melt function created the two columns X1 and X2, which are containing every possible row and column combination, and a third column with the name value, which is containing the corresponding values.
In order to draw a heatmap with the ggplot2 package, we also need to install and load ggplot2:
install.packages("ggplot2") # Install ggplot2 package library("ggplot2") # Load ggplot2 package
Now, we can use the geom_tile function of the ggplot2 package to make a basic heatmap:
ggp <- ggplot(data_melt, aes(X1, X2)) + # Create heatmap with ggplot2 geom_tile(aes(fill = value)) ggp # Print heatmap
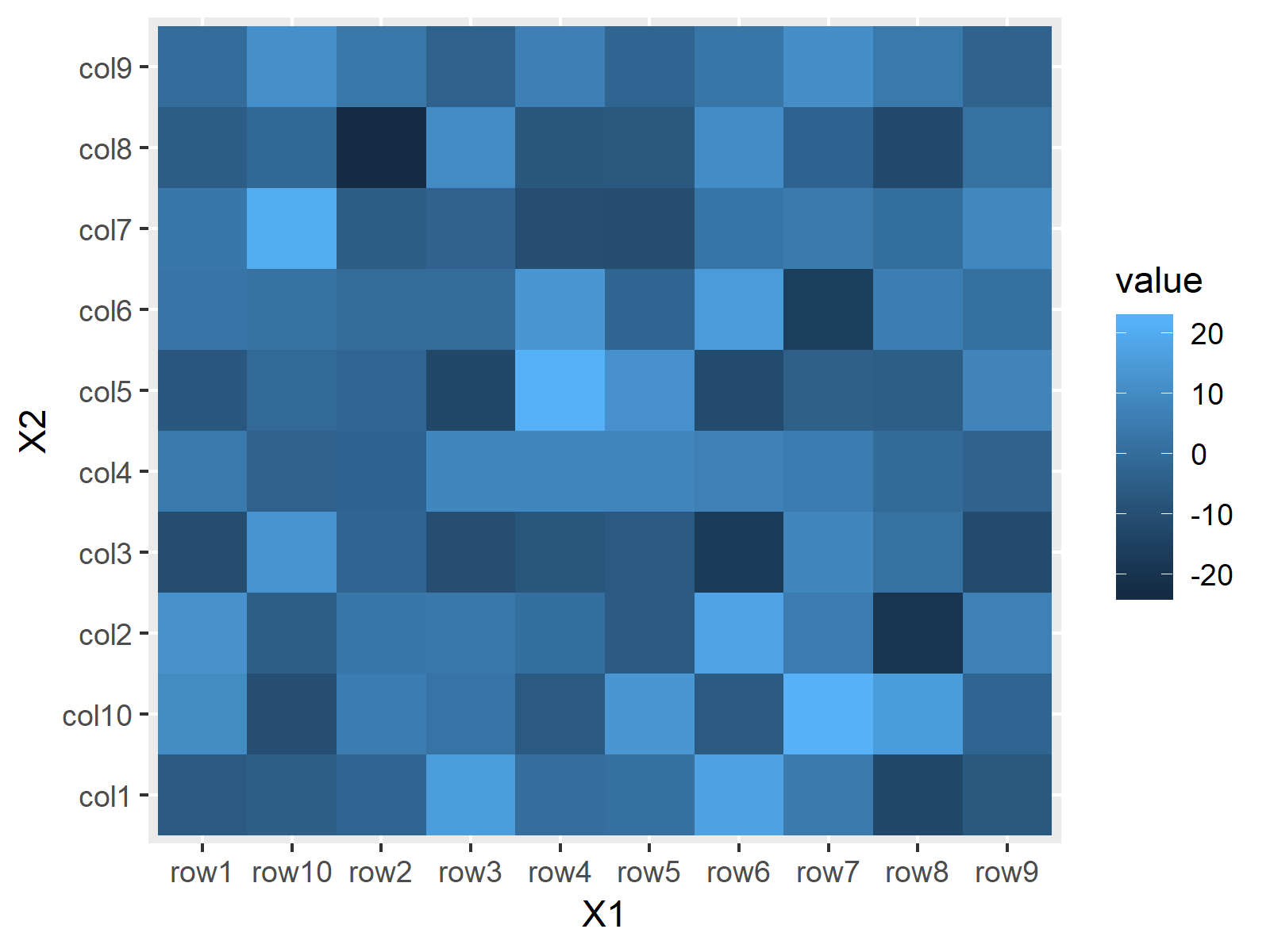
Figure 4: Default Heatmap in ggplot2 Package.
As you can see based on Figure 4, the patter of the heatmap cells is the same as in Base R. However, the general layout is in the typical ggplot2 style.
Of cause, ggplot2 also provides options for the modification of our heatmap. For instance, we can use the scale_fill_gradient function to draw a heatmap with a manual color range:
ggp + scale_fill_gradient(low = "green", high = "black") # Manual colors of heatmap
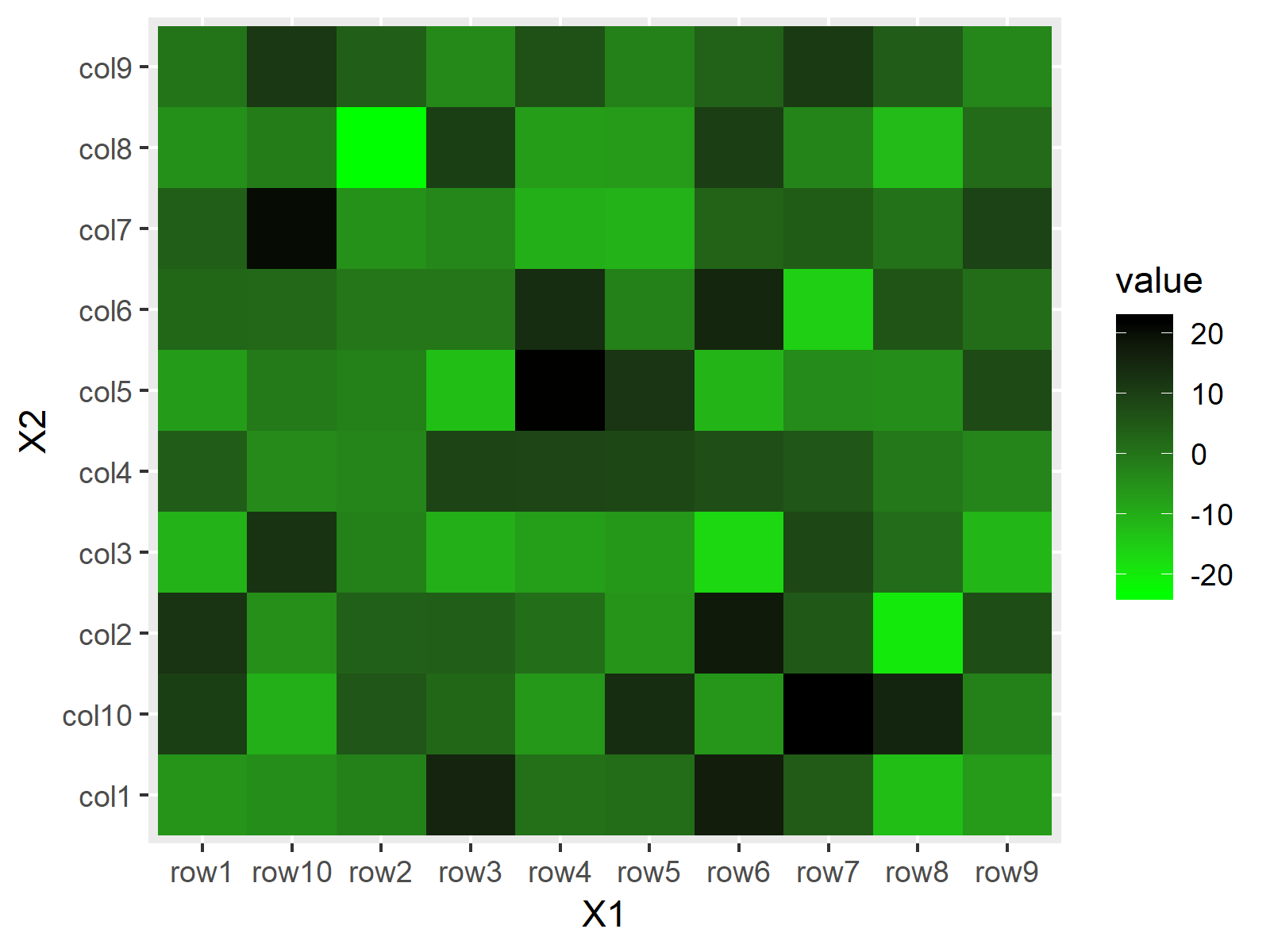
Figure 5: Heatmap with Manual Color Range in ggplot2 Package.
Example 3: Create Heatmap with plot_ly Function [plotly Package]
Another popular package for heatmaps is the plotly package:
install.packages("plotly") # Install plotly package library("plotly") # Load plotly package
The plotly package contains the plot_ly function, which can be used to draw a heatmap by specifying type = “heatmap”:
plot_ly(z = data, type = "heatmap") # Apply plot_ly function
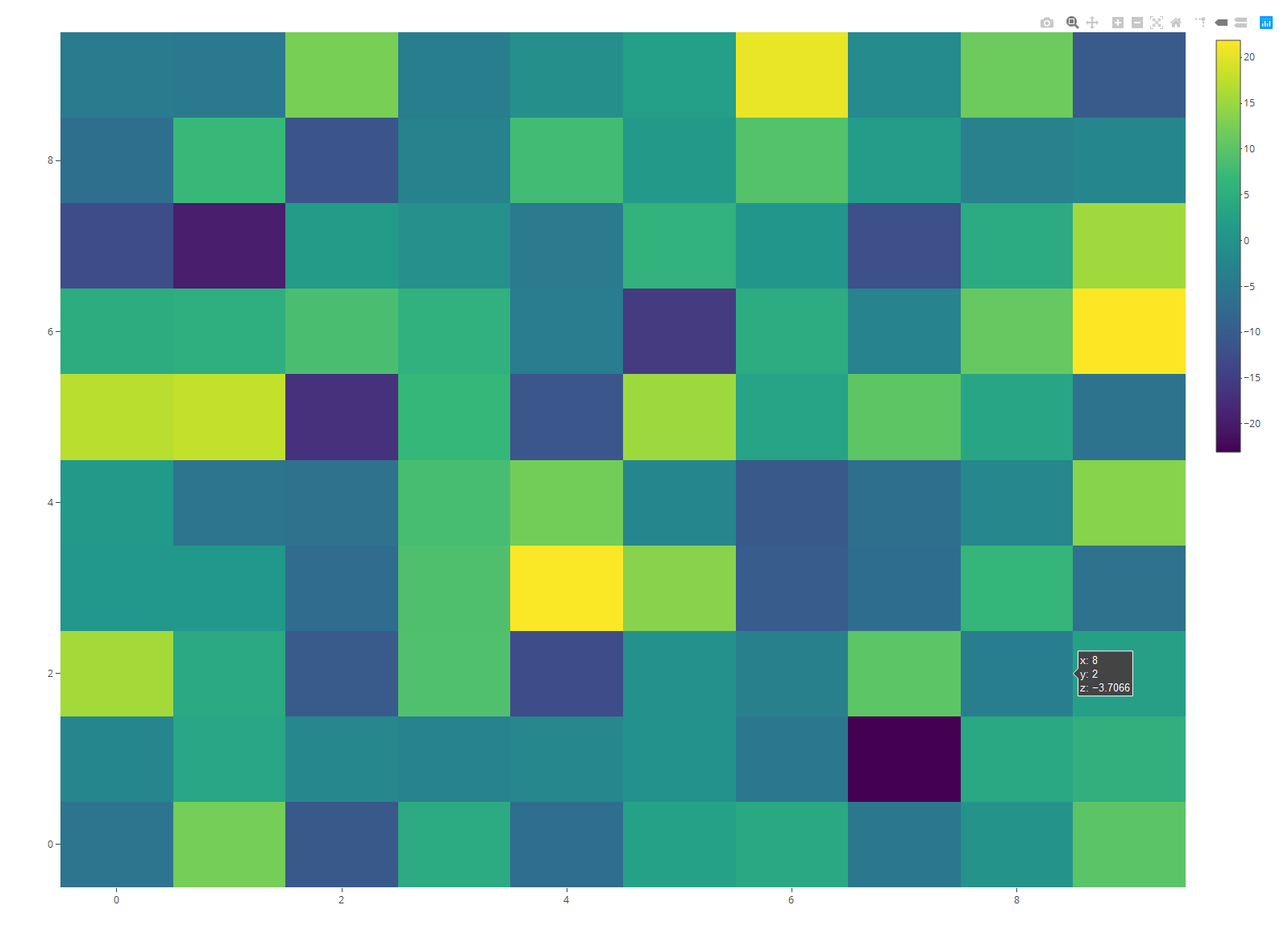
Figure 6: Default Heatmap in plotly Package.
Again, the patter is the same, but the general plot style is different.
The plotly package also provides additional options for the modification of the heatmap. If we want to change the color, we can either specify a color range manually, or we can use some predefined options such as colorscale = “Greys”:
plot_ly(z = data, colorscale = "Greys", type = "heatmap") # Manual colors

Figure 7: Heatmap with Manual Color Range in plotly Package.
As you can see based on Figure 7, the Greys specification created a heatmap in greyscale.
Note that the plotly package show its graphics in the RStudio viewer instead of the RStudio plot window. For that reason you need to export these plots differently.
Also note that there are many other packages for the creation of heatmaps in R available. In my opinion, however, Base R, ggplot2, and plotly provide the best solutions.
Video & Further Resources
I have recently published a video on my YouTube channel, which illustrates the R syntax of this article. You can find the video below:
Also, you might read the other tutorials on my website:
In this R tutorial you learned how to make a heatmap. Don’t hesitate to let me know in the comments below, if you have additional questions.
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.







18 Comments. Leave new
Thanks, Joachim, this is a very helpful introduction to heatmaps.
Thank you Colin, that’s nice to hear!
Hello Joachim, can you please share heatmap example for treatment and control group for RNA SEQ DATA three replicates sample using ggplot2?
For example if I have six columns three repeats of control and three for treatment group.
Hi Umar, please see my comment above.
Hi Umar,
Could you please provide some example data? I’m not sure how your data would look like.
Regards
Joachim
This is really helpful, thank you so much! Regarding exporting plotly heatmaps… just what I was looking for by the way, I love the hover-over function… your blog points to a fairly complicated Stack Overflow thread. In R Studio, I just clicked “export” in the viewer… a bunch of stuff was automatically downloaded and I was able to save a fully-functional HTML version of my heatmap. Kinderleicht!
Thanks Don Pinkerton, I’m glad that you liked the tutorial! Also, thanks a lot for sharing your way of exporting plotly graphs. I think the thread on Stack Overflow is better if you want to automatize the process. However, for a single export your solution is definitely much simpler to apply. Regards, Joachim
This is so well done. Saved me a lot of headache and time
Thanks a lot Patrick, I’m very glad to hear that! 🙂
Thank you Joachi, for a detailed explanation!!
You are very welcome Shandry, glad you liked it! 🙂
Nice work Joachim !!
Thanks a lot Konstantinos, glad to hear that! 🙂
Hi Joachim,
Thank you for your exploratory lecture on Heatmaps.
I want to ask can you share how to split one cell of heatmap into two that shows two different variables.
Following is question that I want to ask
https://stackoverflow.com/questions/65887187/how-to-display-two-categorical-variables-on-one-tile-of-a-heatmap-triangle-til
Hi Farhan,
Thanks for the kind comment! I’ve spent some time to figure out how that works, and to be honest, I also didn’t find a satisfying solution.
This thread on Stack Overflow seems to be promising, but it is based on categorical data, and I assume you are dealing with numerical data.
I have recently created a Facebook discussion group where people can ask questions about R programming and statistics. Could you post your question there? This way, others can contribute/read as well, and maybe somebody in the group knows a good solution for this.
Regards,
Joachim
Hello ,
Thanks alot for this exploratory lecture on Heatmap but i have a question i would like to draw heat map but by using Log2Fc value for two groups how i can do it
Hello Elseedy,
I haven’t used it before, but I found this solution. Would you like to implement something like this?
Regards,
Cansu