R Error in lm.fit(x, y, offset, singular.ok, …) : NA/NaN/Inf in ‘x’ (2 Examples)
In this R tutorial you’ll learn how to deal with the “Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : NA/NaN/Inf in ‘x'”.
The tutorial is structured as follows:
Let’s get started!
Example 1: Data Contains NA, Inf & NaN
The first step is to construct some data that we can use in the following example:
set.seed(52389374) # Create example data data <- data.frame(y = rnorm(100), x = c(NA, Inf, NaN, rnorm(97))) head(data) # Head of example data
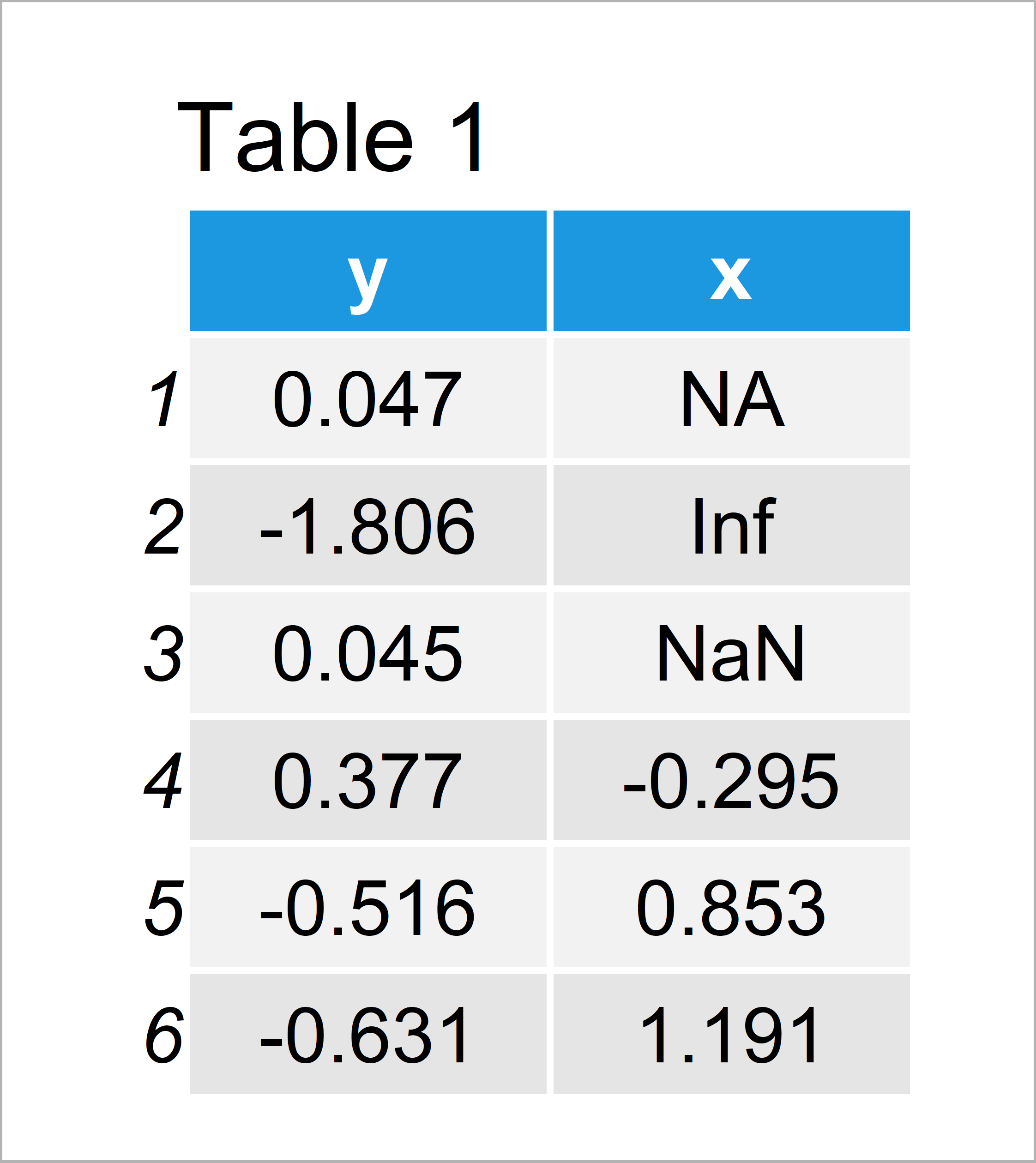
As you can see based on Table 1, our example data is a data frame consisting of 100 rows and two columns.
Based on these data, we can replicate the “Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : NA/NaN/Inf in ‘x'” in the R programming language.
Let’s assume that we want to estimate a linear model based on our data. Then, we typically would apply the lm function as shown below:
lm(y ~ x, data) # Try to apply lm function # Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, ...) : # NA/NaN/Inf in 'x'
Unfortunately, the RStudio console returns the message “Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : NA/NaN/Inf in ‘x'”.
The reason for this is that our data contains NaN and Inf values. In contrast to NA values, these kinds of values cannot be handled by the lm function.
So how can we solve this problem?
To achieve this, we have to replace the NaN and Inf values in our data frame:
data_new <- data # Duplicate data data_new[is.na(data_new) | data_new == "Inf"] <- NA # Replace NaN & Inf with NA
The previous R programming syntax has created a new data frame called data_new that does contain NA values instead of NaN and Inf.
Now, we can apply the lm function to this new data frame:
lm(y ~ x, data_new) # Properly apply lm function # Call: # lm(formula = y ~ x, data = data_new) # # Coefficients: # (Intercept) x # -0.043774 -0.001974
Works fine!
Example 2: Wrong Target Variable in Linear Regression Model
Another reason why the error message “Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : NA/NaN/Inf in ‘x'” occurs is that the target and predictor variables in the lm() function are not specified properly.
Let’s create another example data frame to illustrate that in practice:
set.seed(3334568) # Create example data data2 <- data.frame(x = LETTERS[1:3], y = runif(90)) head(data2) # Head of example data # x y # 1 A 0.47224122 # 2 B 0.14032087 # 3 C 0.15323529 # 4 A 0.08266449 # 5 B 0.10149550 # 6 C 0.68558516
Our data frame contains two variables. The variable y is our outcome, and the variable x is our predictor.
Let’s try to estimate a linear regression model:
my_mod1 <- lm(x ~ y, data2) # Try to estimate model # Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, ...) : # NA/NaN/Inf in 'y' # In addition: Warning message: # In storage.mode(v) <- "double" : NAs introduced by coercion
As you can see, the previous R code has returned an error and a warning message.
The reason for this is that we have specified our dependent and independent variables in the wrong order, i.e. we have tried to use the character variable x as target variable.
Let’s fix this:
my_mod2 <- lm(y ~ x, data2) # Properly estimate model
The previous R code has specified y as the target variable (i.e. on the left side of the ~). This works fine without any error messages.
Video & Further Resources
Do you need more information on the R programming codes of this tutorial? Then I recommend watching the following video of my YouTube channel. In the video, I show the R code of this article in RStudio.
Furthermore, you may read the related tutorials which I have published on my website. You can find a selection of articles on topics such as vectors, coding errors, and missing data below:
- Replace Inf with NA in Vector & Data Frame
- Help – Error in if (NA) { : missing value where TRUE/FALSE needed
- Errors & Warnings in R
- R Programming Tutorials
In this R programming tutorial you have learned how to get rid of the “Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : NA/NaN/Inf in ‘x'”. Let me know in the comments below, if you have further questions. Furthermore, please subscribe to my email newsletter in order to get updates on new tutorials.
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.







16 Comments. Leave new
I have applied this technique but was unsuccessful. My dataset as below. There is no NA/NaN/Inf value.
‘data.frame’: 300 obs. of 6 variables:
$ X : int 1 2 3 4 5 6 7 8 9 10 …
$ Age : num 49.5 42.3 59.4 46.2 26.5 …
$ Sex : chr “M” “F” “M” “F” …
$ Height : num 178 165 172 166 164 …
$ ReactionTime : num 501 415 644 371 508 …
$ AGE_GROUP : chr “40-70” “40-70” “40-70” “40-70” …
lm(AGE_GROUP ~ ReactionTime, dataset)
Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) : NA/NaN/Inf in ‘y’
Any help would be much appreciated. Thanks.
Hey Kafil,
Thanks a lot for your question. Indeed, this reason for the error message has not been covered in the tutorial yet.
I have restructured the tutorial, to include your scenario as well. Please have a look at Example 2 for more details.
In short: I think you have specified your variables in the wrong order (i.e. AGE_GROUP and ReactionTime are exchanged).
Have a look at the reproducible example below:
I hope that helps!
Joachim
My target column has no NA’s but still I am getting the above error and I have selected the correct target variable. please suggest some solution.
Hey Preeti,
Could you illustrate the structure of your data and share your code?
Regards,
Joachim
Hi Joachim,
I had this same warning but my situation doesn’t seem to fit either of your examples.
My code was
reg1_log <- lm(log(sales_MSK) ~ log(price_MSK),
data = da_msk)
my data looks like this :
week store month price_MSK price_LT price_SP store_prom leaflet_ad store_prom_SP leaflet_ad_SP store_prom_LT sales_MSK
1 1 1 2 17.57 19.26 14.54 0 0 0 0 1 2
2 2 1 2 17.57 19.26 14.54 0 0 0 0 0 3
3 3 1 2 17.57 19.26 14.54 0 0 0 0 0 4
4 4 1 2 17.57 19.26 15.29 0 1 0 0 0 6
5 5 1 3 17.57 19.26 15.29 0 0 0 0 0 2
6 6 1 3 17.57 19.26 15.29 1 1 0 0 0 10
7 7 1 3 17.57 19.26 15.29 0 0 0 0 0 3
8 8 1 3 17.57 19.26 15.29 0 0 0 0 0 5
9 9 1 4 17.57 19.26 15.29 0 0 0 0 0 2
10 10 1 4 17.57 19.26 15.29 0 0 0 0 0 3
Hi Jean,
I have tried to reproduce your problem, but with your provided data it works fine for me. Have a look at the example code below:
Have you done anything different as I did?
Regards,
Joachim
Hi Joachim,
Thanks so much for you kind reply. I figured out in the end that in some years the sales was zero and that caused the problem. I ended up use sales+1 instead of sales to solve the problem.
Thanks again for getting back to me 🙂
Ah I see, glad you found a solution! 🙂
Regards,
Joachim
Hi Joachim,
I’m running the following for two datasets, one works fine, but the other I get the error message:
Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) :
NA/NaN/Inf in ‘x
Code:
Group_of_Models %
ungroup()%>%
select(-Biomass, -Dataset, -Month, -Year, -DOY, -Station_Number, -Depth)%>%
map(~lm(log(dataset$Biomass) ~ .x, data = dataset))
Both datasets do not contain Inf values, only NaNs, however, this error does not appear for one of the datasets. Even when I remove the NaNs, the models that contain log(.x) are ignored and not run. I’ve honestly tried pretty much every solution I could find. Not sure what else to do, so any help is greatly appreciated! It’s important to note that this same code was running just fine three weeks ago. I’ve also tried running in different computers and the same message appears.
Best!
Hey Lívia,
Please excuse the late response. I was on a long holiday so unfortunately I wasn’t able to reply sooner. Still need help with your code?
Regards,
Joachim
Hi Joachim,
I’m running the datasets, but I get the error message:
Error in lm.fit(x = ys.lagged, y = yendog) : NA/NaN/Inf in ‘x’
In addition: Warning messages:
1: In lm.fit(x = ys.lagged, y = yendog) : NAs introduced by coercion
2: In lm.fit(x = ys.lagged, y = yendog) : NAs introduced by coercion
Code:
v1 <-VARselect(data,lag.max=10)
My data looks like this
date BVH PVI BIC PGI BMI VND IVS PSI AGR SHS VCB ACB CTG
1 2012-01-03 52.71 45.41 30.11 2.19 3.04 0.90 3.86 3.1 4.05 0.71 8.38 4.54 6.99
2 2012-01-04 52.15 45.03 28.05 2.16 3.18 0.87 3.61 3.1 3.87 0.67 8.18 4.61 7.32
3 2012-01-05 51.30 44.57 29.21 2.16 3.11 0.82 3.61 3.1 3.96 0.65 8.14 4.71 7.28
4 2012-01-06 50.46 44.29 28.88 2.23 3.08 0.78 3.61 3.0 3.96 0.63 7.98 4.68 7.41
5 2012-01-09 50.64 44.29 28.05 2.33 2.93 0.77 3.69 3.1 3.96 0.65 8.22 4.66 7.45
6 2012-01-10 50.64 44.29 28.88 2.33 2.93 0.82 3.43 3.3 4.14 0.69 8.30 4.68 7.69
7 2012-01-11 50.83 44.19 28.88 2.29 3.04 0.81 3.43 3.3 4.23 0.69 8.30 4.66 7.69
8 2012-01-12 49.43 43.36 28.96 2.26 3.04 0.78 3.52 3.3 4.41 0.67 8.46 4.66 8.06
9 2012-01-13 48.39 42.98 28.79 2.29 2.93 0.81 3.35 3.3 4.59 0.69 8.50 4.68 8.43
10 2012-01-16 48.68 42.89 28.75 2.39 3.04 0.85 3.43 3.5 4.77 0.71 8.66 4.73 8.48
11 2012-01-17 49.05 43.08 29.29 2.49 3.04 0.82 3.61 3.4 4.59 0.67 8.62 4.78 8.89
12 2012-01-18 49.24 43.08 29.70 2.43 2.97 0.83 3.61 3.4 4.41 0.71 8.74 4.80 9.05
13 2012-01-19 47.92 42.80 29.70 2.53 2.93 0.86 3.61 3.5 4.59 0.74 9.15 4.88 9.50
14 2012-01-20 48.39 42.70 30.44 2.59 3.08 0.87 3.78 3.4 4.77 0.69 9.31 4.90 9.46
15 2012-01-30 49.71 43.91 31.68 2.66 3.18 0.91 3.95 3.4 4.95 0.71 9.75 4.97 9.75
16 2012-01-31 50.46 44.10 31.68 2.66 3.04 0.92 3.86 3.7 4.77 0.74 9.83 5.07 9.46
17 2012-02-01 51.02 45.87 31.85 2.73 3.04 0.95 3.52 3.6 4.77 0.71 9.59 5.04 9.59
18 2012-02-02 52.05 45.22 32.42 2.79 3.11 0.99 3.61 3.8 4.95 0.76 10.03 5.21 10.04
19 2012-02-03 51.96 45.31 32.92 2.79 3.18 0.98 3.86 3.6 5.04 0.76 9.87 5.11 10.04
20 2012-02-06 52.43 45.41 35.06 2.66 3.15 1.01 3.95 3.6 4.95 0.74 9.79 5.11 9.55
21 2012-02-07 51.96 45.50 34.90 2.73 3.01 1.04 3.86 3.6 4.77 0.76 9.87 5.35 10.00
22 2012-02-08 51.68 45.69 33.83 2.66 3.04 1.04 4.21 3.6 4.95 0.80 10.28 5.47 10.25
23 2012-02-09 50.83 45.03 35.06 2.63 3.08 1.00 3.95 3.5 4.95 0.80 10.15 5.59 10.20
24 2012-02-10 51.39 45.03 35.06 2.59 3.08 0.96 3.95 3.4 4.77 0.78 9.91 5.55 9.87
25 2012-02-13 53.83 45.41 33.08 2.56 3.08 0.91 3.86 3.4 4.59 0.74 9.75 5.31 9.38
26 2012-02-14 53.36 45.59 33.00 2.66 3.04 0.96 3.86 3.4 4.77 0.76 10.15 5.21 9.67
27 2012-02-15 53.74 45.50 33.00 2.56 3.08 0.92 3.61 3.4 4.59 0.71 10.07 5.02 9.42
28 2012-02-16 53.65 45.59 31.35 2.53 3.08 0.95 3.78 3.3 4.68 0.71 10.07 5.09 9.46
29 2012-02-17 52.80 45.59 33.50 2.59 3.08 1.00 4.12 3.6 4.77 0.74 10.28 5.16 9.63
30 2012-02-20 54.02 46.34 33.33 2.56 3.18 1.04 4.29 3.7 4.95 0.78 10.76 5.47 10.08
31 2012-02-21 55.33 46.34 32.92 2.63 3.22 1.07 4.29 3.6 4.77 0.80 10.84 5.40 10.12
32 2012-02-22 55.33 46.43 33.00 2.66 3.29 1.16 4.55 3.9 4.95 0.85 11.32 5.55 10.29
33 2012-02-23 55.43 46.43 30.86 2.73 3.33 1.19 4.64 4.0 5.13 0.87 11.48 5.57 10.37
34 2012-02-24 56.08 47.92 32.14 2.66 3.26 1.18 4.72 4.0 5.22 0.89 11.82 5.47 10.16
35 2012-02-27 55.33 47.36 31.43 2.59 3.33 1.23 5.24 4.3 5.40 0.96 11.91 5.64 10.37
36 2012-02-28 55.33 47.55 31.43 2.59 3.33 1.13 5.58 4.2 5.49 0.94 11.57 5.50 10.33
37 2012-02-29 55.99 46.71 31.35 2.56 3.43 1.18 5.07 4.5 5.58 1.07 11.44 5.62 9.96
38 2012-03-01 55.24 46.15 30.53 2.63 3.58 1.18 5.67 4.6 5.58 1.09 11.95 5.93 10.41
39 2012-03-02 54.77 45.69 30.40 2.59 3.72 1.23 5.50 4.7 5.85 1.16 12.20 6.24 10.78
40 2012-03-05 52.90 46.25 30.40 2.69 3.90 1.30 6.27 5.0 6.12 1.23 12.79 6.64 11.31
41 2012-03-06 53.83 47.09 30.07 2.69 3.72 1.32 6.01 5.3 6.39 1.27 12.16 6.45 11.19
42 2012-03-07 53.46 46.34 29.70 2.69 3.54 1.33 6.53 5.6 6.66 1.32 12.16 6.24 11.19
43 2012-03-08 51.68 46.25 28.88 2.66 3.54 1.23 6.01 5.1 6.93 1.27 11.57 5.90 10.66
44 2012-03-09 51.68 46.34 27.06 2.66 3.40 1.24 5.84 5.3 6.66 1.23 11.74 5.88 10.49
45 2012-03-12 52.71 46.25 28.88 2.66 3.33 1.17 5.75 4.8 6.39 1.18 11.27 5.69 10.00
46 2012-03-13 52.71 46.53 28.67 2.69 3.47 1.24 5.84 4.9 6.39 1.21 11.48 5.88 10.16
47 2012-03-14 52.61 46.34 28.88 2.69 3.51 1.19 5.50 4.7 6.12 1.12 11.65 5.86 10.29
The dataset do not contain Inf/NaNs values, and I cannot go through.
Regards,
Hello,
Sorry for the late response. I am not sure if you could fix it. But I can not match the variables you use in the lm() function and the dataset. Am I missing something?
Regards,
Cansu
Hi Joachim
thank you for your explanations; but I have the same problem and I don’t know how to solve it. In my data I don’t have Nan or Inf but when I fit the linear model I obtain the error:
Error in lm.fit(x, y, offset = offset, singular.ok = singular.ok, …) :
NA/NaN/Inf dans ‘y’
In addition: Warning message:
In log(Y) : NaNs produced
Here are my datas:
2 3 4 5 6
[1,] 23451137.9 8696791.6 8794749.66 3876954.62 3229763.5
[2,] 8114722.1 1518537.1 -225003.67 2624349.38 740171.5
[3,] 3410104.6 10858244.2 -331044.40 6060053.81 -9960230.9
[4,] 34535804.0 -3682029.3 3498229.50 1491422.65 1392363.0
[5,] 33614304.4 24741513.1 3868686.16 115029.49 -249767.4
[6,] 3703572.5 101049649.2 -10639025.63 11923.38 9598247.7
[7,] 0.0 0.0 0.00 0.00 0.0
[8,] 2033708.8 12632522.9 1520242.78 262761.42 -616142.7
[9,] 63811431.4 2224806.0 -175592.51 1007724.46 1192133.8
[10,] 1146828.3 1231986.3 61848.07 -128514.73 -126501.9
[11,] 1014042.3 219266.4 -2018282.41 1462843.94 NA
[12,] 10526206.2 -882072.1 -646326.31 NA NA
[13,] 5654361.5 -695112.3 NA NA NA
[14,] 389327.4 NA NA NA NA
[15,] NA NA NA NA NA
Hello Denise,
You may receive this error due to the 15th row of your data frame. The fact that you have only NAs might be one reason that you get this error. Could you please share the code of your model? Also, the second error may imply that you have an undefined logarithmic function, log(0) is undefined, for instance. This may be due to the row of 0s in your data frame. But since I don’t know your model, it is hard to test them. You can also try to test it by yourself by removing those rows and see if the model runs or not.
Regards,
Cansu
Hi Cansu,
thank youi for your answer but I’ve tried it to no avail always.
Please find below the code of my model:
PAID<-read_excel("aviation.xlsx",na="NA", col_names = TRUE)
PAID<-as.matrix(PAID)
PAID
nc<-ncol(PAID)
nl<-nrow(PAID)
ligne <- rep(1:nl, each=nc); colonne <- rep(1:nc, nl)
INC <- PAID
INC[,2:6] <- PAID[,2:6]-PAID[,1:5]
Y <- as.vector(abs(INC))
lig <- as.factor(ligne)
col <- as.factor(colonne)
reg <- lm(log(Y)~col+lig) ## this is the line from where I got the error
summary(reg)
log(Y).
My database PAID is given by :
1571462.24000000 25022600.1500000 33719391.7800000 42514141.4400000 46391096.0600000 49620859.560000 53295824.8800000 80419958.1600
[2,] 335705.34000000 8450427.3900000 9968964.5400000 9743960.8700000 12368310.2500000 13108481.740000 13890889.7900000 16162550.7400
[3,] 1831770.00000000 5241874.5900000 16100118.8400000 15769074.4400000 21829128.2500000 11868897.390000 12963201.1300000 11465508.8900
[4,] 1851440.00000000 36387243.9800000 32705214.6900000 36203444.1900000 37694866.8400000 39087229.880000 52584783.7800000 53085469.9700
[5,] 17188626.68000000 50802931.0700000 75544444.2000000 79413130.3600000 79528159.8500000 79278392.440000 79278392.4400000 79278392.4400
[6,] 1330865.16000000 5034437.6400000 106084086.8700000 95445061.2400000 95456984.6200000 105055232.310000 107510600.2000000 106438365.9300
[7,] 0.00010000 0.0001000 0.0001000 0.0001000 0.0001000 0.000100 0.0001000 0.0001
[8,] 4114806.51000000 6148515.3000000 18781038.2400000 20301281.0200000 20564042.4400000 19947899.720000 19871333.1959128 20489280.7300
[9,] 39258492.66000000 103069924.0100000 105294729.9700000 105119137.4600000 106126861.9200000 107318995.677048 106923891.8529951 NA
[10,] 107971.68000000 1254799.9900000 2486786.2800000 2548634.3500000 2420119.6163248 2293617.690000 NA NA
[11,] 28537423.17000000 29551465.4900000 29770731.8500000 27752449.4372951 29215293.3800000 NA NA NA
[12,] 4711641.37000000 15237847.5700000 14355775.4632837 13709449.1500000 NA NA NA NA
[13,] 4529909.17000000 10184270.6551324 9489158.3400000 NA NA NA NA NA
[14,] 6595109.42138589 6984436.8000000 NA NA NA NA NA NA
[15,] 11140590.55000000 NA NA NA NA NA NA NA
9 10 11 12 13 14 15
[1,] 82447132.160000 82447132.1600 82447132.1600000 82447132.1600000 82447132.1600000 82447132.16 82447132.16
[2,] 22726503.560000 20121944.5100 20135395.3700000 20124956.3900000 20073809.3559984 20073873.43 NA
[3,] 10107427.130000 12169782.3600 10229675.2600000 10706614.9305173 11100890.8600000 NA NA
[4,] 53296993.680000 53442127.8100 55752767.6476081 61868552.7700000 NA NA NA
[5,] 79278392.440000 85159239.8802 85159239.8802000 NA NA NA NA
[6,] 106383659.007709 106753743.5800 NA NA NA NA NA
[7,] 0.000100 NA NA NA NA NA NA
[8,] NA NA NA NA NA NA NA
[9,] NA NA NA NA NA NA NA
[10,] NA NA NA NA NA NA NA
[11,] NA NA NA NA NA NA NA
[12,] NA NA NA NA NA NA NA
[13,] NA NA NA NA NA NA NA
[14,] NA NA NA NA NA NA NA
[15,] NA NA NA NA NA NA NA
Thanks!!!
Hello again,
So you say that you tried to remove NA rows, and the model still gives an error, right? If not, could you please try it out? Sorry, I am not familiar with working with matrices for modeling. So it looked to me strange how you don’t use the columns of the dataset as the independent and dependent variables. But I suspect that it is due to the uncomputable log(Y) values, as early told.
Regards,
Cansu