Sum of Two or Multiple Data Frame Columns in R (2 Examples)
In this article you’ll learn how to compute the sum across two or more columns of a data frame in the R programming language.
Table of contents:
Let’s dive into it!
Example Data
The first step is to define some example data:
data <- data.frame(x1 = 1:5, # Create data frame x2 = c(3, 1, 7, 4, 4), x3 = 5:9, x4 = c(7, 4, 6, 4, 9)) data # Print data frame
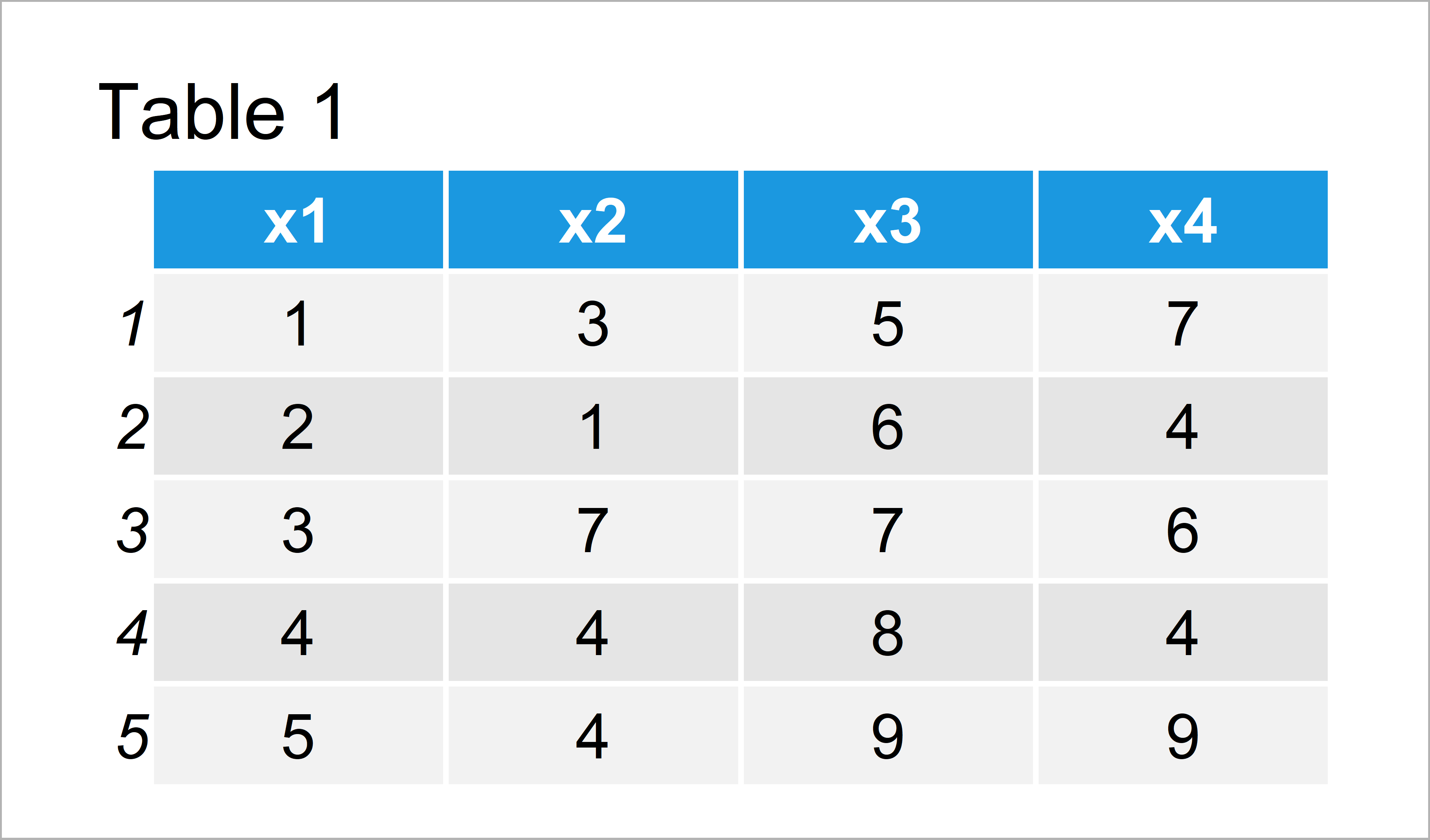
As you can see based on Table 1, our example data is a data frame consisting of five rows and four columns. All the variables are numeric.
Example 1: Calculate Sum of Two Columns Using + Operator
In this example, I’ll explain how to get the sum across two columns of our data frame.
For this, we can use the + and the $ operators as shown below:
data$x1 + data$x2 # Sum of two columns # [1] 4 3 10 8 9
After executing the previous R code, the result is shown in the RStudio console.
Example 2: Calculate Sum of Multiple Columns Using rowSums() & c() Functions
It is also possible to return the sum of more than two variables. To efficiently calculate the sum of the rows of a data frame subset, we can use the rowSums function as shown below:
rowSums(data[ , c("x1", "x2", "x4")]) # Sum of multiple columns # [1] 11 7 16 12 18
The result of the addition of the variables x1, x2, and x4 is shown in the RStudio console.
Video, Further Resources & Summary
Do you want to learn more about sums and data frames in R? Then I recommend having a look at the following video of my YouTube channel. I show the R code of this tutorial in the video:
Also, you might read the other articles on this website.
- Sums of Rows & Columns in Data Frame or Matrix
- Sum Across Multiple Rows & Columns Using dplyr Package
- The R Programming Language
Summary: In this article, I have explained how to calculate the sum of data frame variables in the R programming language. If you have additional questions and/or comments, let me know in the comments section. Furthermore, don’t forget to subscribe to my email newsletter for regular updates on the newest tutorials.






