glm.fit Warning Messages in R: algorithm didn’t converge & probabilities 0/1
In this R tutorial you’ll learn how to handle the glm.fit warning: “algorithm did not converge and fitted probabilities numerically 0 or 1”.
The article is structured as follows:
Let’s dive right in.
Construction of Example Data
The first step is to create some data that we can use in the following examples.
set.seed(6523987) # Create example data x <- rnorm(100) y <- rep(1, 100) y[x < 0] <- 0 data <- data.frame(x, y) head(data) # Head of example data
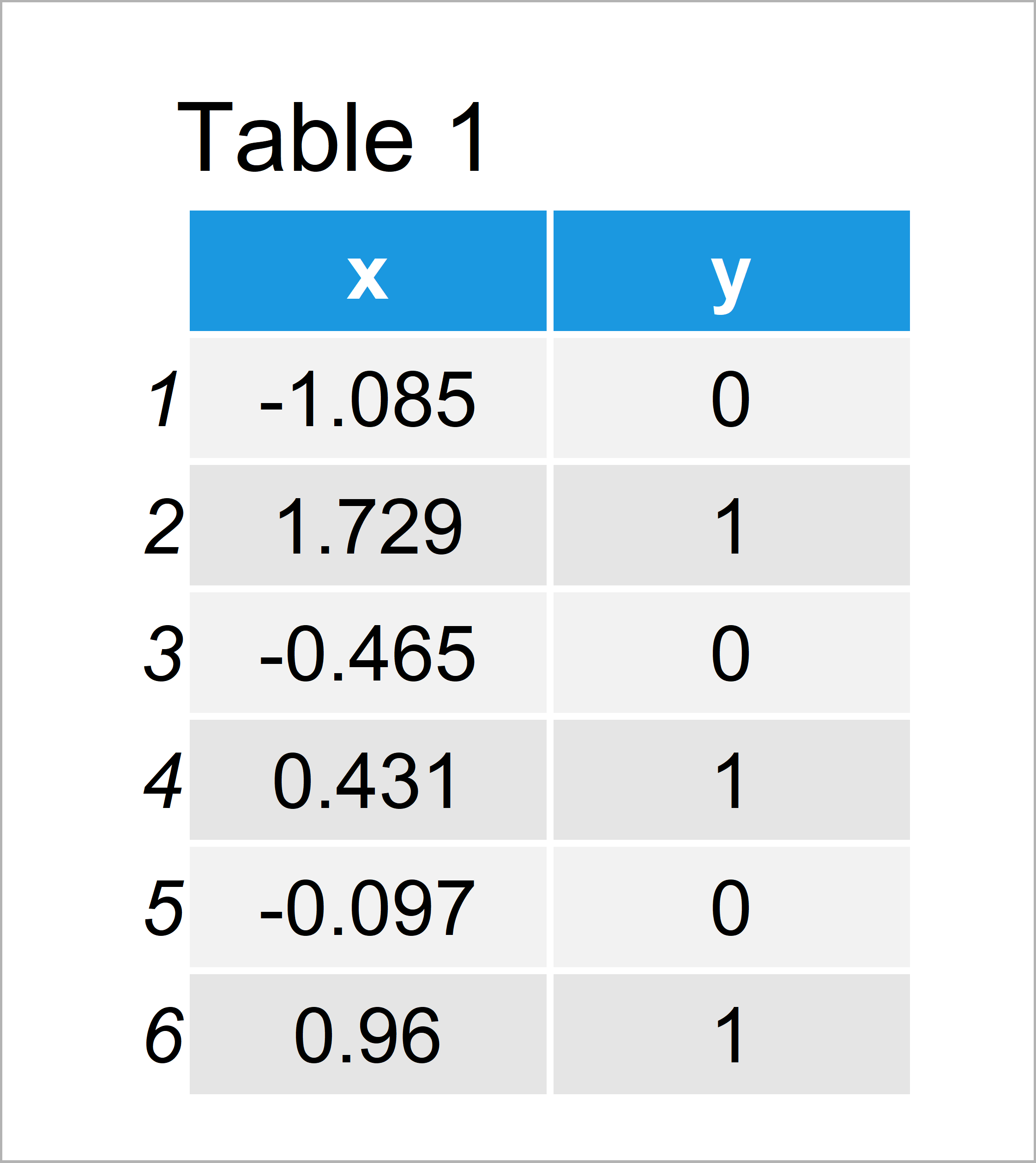
Table 1 shows that our example data consists of 100 rows and two columns x and y.
Example: Reproduce the glm.fit Warning Message – algorithm did not converge fitted probabilities 0 or 1
The following syntax explains why the glm.fit warning: “algorithm did not converge and fitted probabilities numerically 0 or 1” occurs when fitting regression models in the R programming language.
Let’s assume that we want to estimate a binomial model using the glm function. Then we would typically specify our R code as shown below:
glm(y ~ x, data, family = "binomial") # Perfect predictions # Warning messages: # 1: glm.fit: algorithm did not converge # 2: glm.fit: fitted probabilities numerically 0 or 1 occurred
Unfortunately, the glm.fit warning: “algorithm did not converge and fitted probabilities numerically 0 or 1” appears.
The reason for this is that the variable x perfectly predicts the variable y.
You can see that when you go back to the data creation process at the beginning of this tutorial. I have specified that y is always 0 when x is smaller than 0. The model hence perfectly predicts which y values are 0 and which y values are 1.
We can illustrate that by adding some random noise to our data:
data_new <- data # Add randomness to data data_new$x <- data_new$x + rnorm(100)
Let’s run a glm model with our new data:
glm(y ~ x, data_new, family = "binomial") # No warning message # # Call: glm(formula = y ~ x, family = "binomial", data = data_new) # # Coefficients: # (Intercept) x # -0.1049 1.4463 # # Degrees of Freedom: 99 Total (i.e. Null); 98 Residual # Null Deviance: 138 # Residual Deviance: 90.85 AIC: 94.85
This time we don’t get a warning message in the RStudio console.
So how can we solve this problem?
This depends heavily on the structure of the specific data set you are using. However, I advise to have a look at the pairwise correlations of the variables in your data to see if some of these correlations are unnaturally high.
It often happens that variables are included to a model by accident. Try to identify these variables and run your model again without these variables until the warning message does not appear anymore.
Video, Further Resources & Summary
Have a look at the following video of my YouTube channel. I’m showing the R programming syntax of this post in the video:
The YouTube video will be added soon.
Furthermore, you may want to have a look at the related tutorials on https://statisticsglobe.com/. I have released numerous tutorials already:
In this R tutorial you have learned how to deal with the glm.fit warning: “algorithm did not converge and fitted probabilities numerically 0 or 1”. In case you have additional questions and/or comments, let me know in the comments.
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.






