Convert Date to Day of Week in R (3 Examples) | How to Find the Weekday
In this article, I’ll explain how to find the weekday of a date in R. The tutorial is structured as follows
- Creation of Example Data in R
- Find Weekday with weekdays Function (Example 1)
- Find Weekday with strftime Function (Example 2)
- Find Weekday with as.POSIXlt Function (Example 3)
- Further Resources for the Handling of Dates
So without further ado, let’s dive into the examples…
Create Example Data
Before we can start with the examples, we have to create some example data that contains a date variable:
data <- data.frame(date = as.Date(c("2022-02-11", # Create example data "2010-12-01", "1955-02-08", "2005-01-01"))) data # Print example data to console
Table 1: Example Data Frame with Dates.
As you can see based on Table 1, our example data frame consists of a column with dates. Note that these dates were converted to date objects with the as.Date function.
Now let’s move on to the examples, where we will find the weekdays that correspond to our dates…
Example 1: Convert Date to Weekday in R (weekdays Function)
In the first example, we are converting our dates to weekdays with the weekdays function. The function is explicitly designed to find the corresponding day of the week for a date object in R:
data1 <- data # Replicate data for Example 1 data1$weekday <- weekdays(data1$date) # Convert dates to weekdays data1 # Print converted data to console
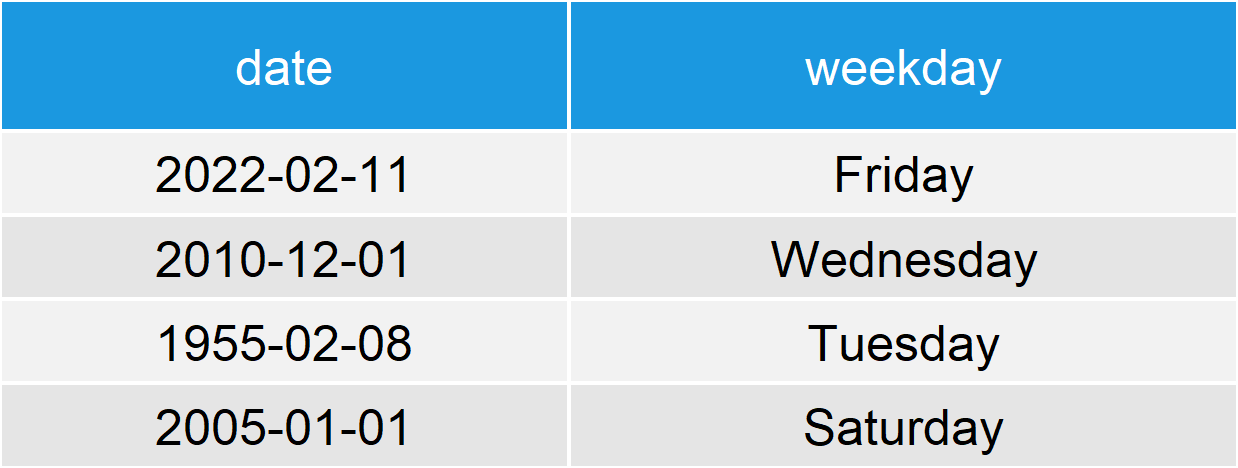
Table 2: Example Data Frame with Dates & Weekdays.
As you can see in Table 2, we have just added a second variable to our data frame which contains the weekday for each of our dates.
Note: We were able to find the weekdays of dates in the past as well as of dates in the future (as in row 1).
The weekdays function is quite simple and easy to use. However, there are many other ways of how to find weekdays and depending on your specific situation one of the following approaches might be better. So keep on reading…
Example 2: Convert Date to Weekday in R (strftime Function)
In the second example, we are using the strftime function:
data2 <- data # Replicate data for Example 2 data2$day <- strftime(data2$date, "%A") data2 # Print converted data to console # date weekday # 2022-02-11 Friday # 2010-12-01 Wednesday # 1955-02-08 Tuesday # 2005-01-01 Saturday
The output is exactly the same as in Example 1.
Looks good, so let’s move on to the next example…
Example 3: Convert Date to Weekday in R (as.POSIXlt Function)
In this example we are going to apply the as.POSIXlt command in order to find the corresponding days of the week:
data3 <- data # Replicate data for Example 3 data3$weekday <- c("Sunday", "Monday", "Tuesday", # Convert dates to weekdays "Wednesday", "Thursday", "Friday", "Saturday")[as.POSIXlt(data3$date)$wday + 1] data3 # Print converted data to console # date weekday # 2022-02-11 Friday # 2010-12-01 Wednesday # 1955-02-08 Tuesday # 2005-01-01 Saturday
Again the same output. However, there is a big advantage of this approach: You can assign a name for each day of the week manually. For instance, this has advantages when your R or RStudio are set to a different language than English. In this case the approaches of Example 1 & 2 will assign the weekdays in a different language. With the approach of Example 3, you can assign any character string you want.
Video & Further Resources
For more detailed information concerning the code of this article, please check out the below video on the Statistics Globe YouTube channel:
Handling dates in R is a difficult topic. For that reason I have listed some further resources about the handling of date objects in the following.
If you want to learn more about date objects in R, I can recommend the following YouTube video of Vincent King:
Also, you might have a look at the following R tutorials of statisticsglobe.com:
- Convert Date to Numeric Time Object in R
- weekdays, months, quarters & julian R Functions
- strptime & strftime R Functions
- as.Date Function in R
- The difftime R Function
- List of Useful R Functions
- The R Programming Language
I hope you found everything you were looking for in this tutorial. Let me know in the comments if you have any further questions. Of course, general feedback is also very welcome!







18 Comments. Leave new
didn’t work
Could you tell me some more details? 😉
For 1st 2 options: :
Warning in install.packages :
package ‘weekdays’ is not available for this version of R
A version of this package for your version of R might be available elsewhere,
see the ideas at
https://cran.r-project.org/doc/manuals/r-patched/R-admin.html#Installing-packages
Warning in install.packages :
package ‘strftime’ is not available for this version of R
A version of this package for your version of R might be available elsewhere,
see the ideas at
https://cran.r-project.org/doc/manuals/r-patched/R-admin.html#Installing-packages
And for POSIXlt – Unable to locate from install or repository also
Hey Ravi,
Which version of R are you using? In case it is an older version, I recommend updating it as explained here: https://www.linkedin.com/pulse/3-methods-update-r-rstudio-windows-mac-woratana-ngarmtrakulchol/
Regards
Joachim
Hello Joachim, I am sorry for bother you.
I am trying the third example you gave above since the code returns the weekdays in spanish and is not working.
I’ve got the follow error:
Error in `$<-.data.frame`(`*tmp*`, weekday, value = c("Tuesday", "Wednesday", :
replacement has 355 rows, data has 410
Could you please help me?
Thanks in advance
Hey Ana,
Could you please share your code and illustrate the structure of your data in some more detail? How are your dates formatted?
Regards,
Joachim
“`{r}
weekday_steps_sleep <- daily_activity_sleep
weekday_steps_sleep$weekday <- weekdays(weekday_steps_sleep$date)
“`
'data.frame': 410 obs. of 19 variables:
$ id : num 1.5e+09 1.5e+09 1.5e+09 1.5e+09 1.5e+09 …
$ date : Date, format: "2016-04-12" "2016-04-13" …
$ totalsteps : num 13162 10735 9762 12669 9705 …
$ totaldistance : num 8.5 6.97 6.28 8.16 6.48 …
$ trackerdistance : num 8.5 6.97 6.28 8.16 6.48 …
$ loggedactivitiesdistance: num 0 0 0 0 0 0 0 0 0 0 …
$ veryactivedistance : num 1.88 1.57 2.14 2.71 3.19 …
$ moderatelyactivedistance: num 0.55 0.69 1.26 0.41 0.78 …
$ lightactivedistance : num 6.06 4.71 2.83 5.04 2.51 …
$ sedentaryactivedistance : num 0 0 0 0 0 0 0 0 0 0 …
$ veryactiveminutes : num 25 21 29 36 38 50 28 19 41 39 …
$ fairlyactiveminutes : num 13 19 34 10 20 31 12 8 21 5 …
$ lightlyactiveminutes : num 328 217 209 221 164 264 205 211 262 238 …
$ sedentaryminutes : num 728 776 726 773 539 775 818 838 732 709 …
$ calories : num 1985 1797 1745 1863 1728 …
$ totalsleeprecords : num 1 2 1 2 1 1 1 1 1 1 …
$ totalminutesasleep : num 327 384 412 340 700 304 360 325 361 430 …
$ totaltimeinbed : num 346 407 442 367 712 320 377 364 384 449 …
$ weekday : chr "martes" "miércoles" "viernes" "sábado" …
“`{r}
daily_activity_sleep <- daily_activity_sleep
daily_activity_sleep1$weekday <- c("Monday", "Tuesday", "Wednesday", "Thursday", "Friday", "Saturday", "Sunday")[as.POSIXlt(daily_activity_sleep$date)$wday]
“`
Error in `$<-.data.frame`(`*tmp*`, weekday, value = c("Tuesday", "Wednesday", :
replacement has 355 rows, data has 410
#Let me know if you need any more details.
Thank you very much
Thanks for the further information Ana.
Do the data frames daily_activity_sleep1 and daily_activity_sleep contain the same number of rows? Why are you not using the same data frame in your last line of code?
Regards,
Joachim
Hello Joachim, “daily_activity_sleep1” has typing error. The correct is “daily_activity_sleep”.
And yes, they have the same number of rows.
Show in New Window
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘http://cran.rstudio.com/bin/macosx/contrib/4.2/tidyverse_1.3.2.tgz’
Content type ‘application/x-gzip’ length 420896 bytes (411 KB)
==================================================
downloaded 411 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//RtmpJVaFZN/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘http://cran.rstudio.com/bin/macosx/contrib/4.2/lubridate_1.8.0.tgz’
Content type ‘application/x-gzip’ length 1483283 bytes (1.4 MB)
==================================================
downloaded 1.4 MB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//RtmpJVaFZN/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘http://cran.rstudio.com/bin/macosx/contrib/4.2/dplyr_1.0.9.tgz’
Content type ‘application/x-gzip’ length 1306382 bytes (1.2 MB)
==================================================
downloaded 1.2 MB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//RtmpJVaFZN/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘http://cran.rstudio.com/bin/macosx/contrib/4.2/ggplot2_3.3.6.tgz’
Content type ‘application/x-gzip’ length 4125157 bytes (3.9 MB)
==================================================
downloaded 3.9 MB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//RtmpJVaFZN/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘http://cran.rstudio.com/bin/macosx/contrib/4.2/tidyr_1.2.0.tgz’
Content type ‘application/x-gzip’ length 1001459 bytes (977 KB)
==================================================
downloaded 977 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//RtmpJVaFZN/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘http://cran.rstudio.com/bin/macosx/contrib/4.2/here_1.0.1.tgz’
Content type ‘application/x-gzip’ length 51881 bytes (50 KB)
==================================================
downloaded 50 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//RtmpJVaFZN/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘http://cran.rstudio.com/bin/macosx/contrib/4.2/skimr_2.1.4.tgz’
Content type ‘application/x-gzip’ length 1215018 bytes (1.2 MB)
==================================================
downloaded 1.2 MB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//RtmpJVaFZN/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘http://cran.rstudio.com/bin/macosx/contrib/4.2/janitor_2.1.0.tgz’
Content type ‘application/x-gzip’ length 245505 bytes (239 KB)
==================================================
downloaded 239 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//RtmpJVaFZN/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘http://cran.rstudio.com/bin/macosx/contrib/4.2/webr_0.1.5.tgz’
Content type ‘application/x-gzip’ length 943493 bytes (921 KB)
==================================================
downloaded 921 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//RtmpJVaFZN/downloaded_packages
Show in New Window
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/tidyverse_1.3.2.tgz’
Content type ‘application/x-gzip’ length 420896 bytes (411 KB)
==================================================
downloaded 411 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/lubridate_1.8.0.tgz’
Content type ‘application/x-gzip’ length 1483283 bytes (1.4 MB)
==================================================
downloaded 1.4 MB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/dplyr_1.0.9.tgz’
Content type ‘application/x-gzip’ length 1306382 bytes (1.2 MB)
==================================================
downloaded 1.2 MB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/ggplot2_3.3.6.tgz’
Content type ‘application/x-gzip’ length 4125157 bytes (3.9 MB)
==================================================
downloaded 3.9 MB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/tidyr_1.2.0.tgz’
Content type ‘application/x-gzip’ length 1001459 bytes (977 KB)
==================================================
downloaded 977 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/here_1.0.1.tgz’
Content type ‘application/x-gzip’ length 51881 bytes (50 KB)
==================================================
downloaded 50 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/skimr_2.1.4.tgz’
Content type ‘application/x-gzip’ length 1215018 bytes (1.2 MB)
==================================================
downloaded 1.2 MB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/janitor_2.1.0.tgz’
Content type ‘application/x-gzip’ length 245505 bytes (239 KB)
==================================================
downloaded 239 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/webr_0.1.5.tgz’
Content type ‘application/x-gzip’ length 943493 bytes (921 KB)
==================================================
downloaded 921 KB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Installing package into ‘/Users/anadoamaral/Library/R/x86_64/4.2/library’
(as ‘lib’ is unspecified)
probando la URL ‘https://cran.rstudio.com/bin/macosx/contrib/4.2/clock_0.6.1.tgz’
Content type ‘application/x-gzip’ length 8540466 bytes (8.1 MB)
==================================================
downloaded 8.1 MB
The downloaded binary packages are in
/var/folders/0z/511yr_m53kg4hg4j_qhzcn900000gp/T//Rtmpd05Wsw/downloaded_packages
Show in New Window
Registered S3 methods overwritten by ‘dbplyr’:
method from
print.tbl_lazy
print.tbl_sql
── Attaching packages ───────────────────────────────────── tidyverse 1.3.2 ──✔ ggplot2 3.3.6 ✔ purrr 0.3.4
✔ tibble 3.1.7 ✔ dplyr 1.0.9
✔ tidyr 1.2.0 ✔ stringr 1.4.0
✔ readr 2.1.2 ✔ forcats 0.5.1── Conflicts ──────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
Attaching package: ‘lubridate’
The following objects are masked from ‘package:base’:
date, intersect, setdiff, union
here() starts at /Users/anadoamaral
Registered S3 methods overwritten by ‘htmltools’:
method from
print.html tools:rstudio
print.shiny.tag tools:rstudio
print.shiny.tag.list tools:rstudio
Attaching package: ‘janitor’
The following objects are masked from ‘package:stats’:
chisq.test, fisher.test
Registered S3 method overwritten by ‘data.table’:
method from
print.data.table
Registered S3 method overwritten by ‘htmlwidgets’:
method from
print.htmlwidget tools:rstudio
Attaching package: ‘clock’
The following object is masked from ‘package:lubridate’:
as_date
Show in New Window
A tibble:22,099 × 4
Id
date
time
StepTotal
1503960366 04/12/2016 12:00:00 AM 373
1503960366 04/12/2016 01:00:00 AM 160
1503960366 04/12/2016 02:00:00 AM 151
1503960366 04/12/2016 03:00:00 AM 0
1503960366 04/12/2016 04:00:00 AM 0
1503960366 04/12/2016 05:00:00 AM 0
1503960366 04/12/2016 06:00:00 AM 0
1503960366 04/12/2016 07:00:00 AM 0
1503960366 04/12/2016 08:00:00 AM 250
1503960366 04/12/2016 09:00:00 AM 1864
…
1-10 of 22,099 rows
Show in New Window
A tibble:6 × 4
Id
date
time
StepTotal
1503960366 04/12/2016 12:00:00 AM 373
1503960366 04/12/2016 01:00:00 AM 160
1503960366 04/12/2016 02:00:00 AM 151
1503960366 04/12/2016 03:00:00 AM 0
1503960366 04/12/2016 04:00:00 AM 0
1503960366 04/12/2016 05:00:00 AM 0
6 rows
Show in New Window
‘data.frame’: 410 obs. of 18 variables:
$ id : num 1.5e+09 1.5e+09 1.5e+09 1.5e+09 1.5e+09 …
$ date : Date, format: “2016-04-12” “2016-04-13” …
$ totalsteps : num 13162 10735 9762 12669 9705 …
$ totaldistance : num 8.5 6.97 6.28 8.16 6.48 …
$ trackerdistance : num 8.5 6.97 6.28 8.16 6.48 …
$ loggedactivitiesdistance: num 0 0 0 0 0 0 0 0 0 0 …
$ veryactivedistance : num 1.88 1.57 2.14 2.71 3.19 …
$ moderatelyactivedistance: num 0.55 0.69 1.26 0.41 0.78 …
$ lightactivedistance : num 6.06 4.71 2.83 5.04 2.51 …
$ sedentaryactivedistance : num 0 0 0 0 0 0 0 0 0 0 …
$ veryactiveminutes : num 25 21 29 36 38 50 28 19 41 39 …
$ fairlyactiveminutes : num 13 19 34 10 20 31 12 8 21 5 …
$ lightlyactiveminutes : num 328 217 209 221 164 264 205 211 262 238 …
$ sedentaryminutes : num 728 776 726 773 539 775 818 838 732 709 …
$ calories : num 1985 1797 1745 1863 1728 …
$ totalsleeprecords : num 1 2 1 2 1 1 1 1 1 1 …
$ totalminutesasleep : num 327 384 412 340 700 304 360 325 361 430 …
$ totaltimeinbed : num 346 407 442 367 712 320 377 364 384 449 …
Thanks once again.
What is returned when you run the following line of code:
Regards,
Joachim
Description:df [6 × 18]
id
date
totalsteps
totaldistance
trackerdistance
1 1503960366 2016-04-12 13162 8.50 8.50
2 1503960366 2016-04-13 10735 6.97 6.97
3 1503960366 2016-04-15 9762 6.28 6.28
4 1503960366 2016-04-16 12669 8.16 8.16
5 1503960366 2016-04-17 9705 6.48 6.48
6 1503960366 2016-04-19 15506 9.88 9.88
Thank you once again for the clarifications Ana! I have noticed that the as.POSIXlt function returns Sunday as the value 0 instead of 7, and for that reason, the code of Example 3 didn’t work properly in your situation.
I’m very sorry for this. I don’t know whether the output of the as.POSIXlt function has changed over time (the tutorial is quite old), or if I’ve made a mistake when I created the tutorial.
However, the example code below should work for your data:
I hope that helps!
Joachim
Hi Joahim, I found a solution!
“`{r}
Sys.setlocale(“LC_TIME”, “C”)
“`
Now, I am just struggling to change the column of Weekdays from Chr to date!
Glad you found a solution!
I don’t know whether it’s possible to convert a weekday character string to the Date class. Is this necessary in your case? What do you want to do with the weekdays column?
Hey Joachim, I really appreciate your kindness. I will reference you in my project!
I need to plot the dataset which contains this weekday column.
“`{r}
ggplot(weekday_steps_sleep) +
geom_col(aes(weekday, daily_steps), fill = “#006699”) +
geom_hline(yintercept = 7500) +
labs(title = “Daily steps per weekday”, x= “”, y = “”) +
theme(axis.text.x = element_text(angle = 45,vjust = 0.5, hjust = 1))
“`
Since the weekday column is character string, my graph read the weekdays in alphabetic order and not by weekdays.
Thank you very much for the reference Ana, that’s very kind! 🙂
You can reorder the bars of your barplot as explained here. There’s no need to convert your weekdays to the Date class.
Regards,
Joachim
It worked!!! Vielen vielen dank!!!
That is great to hear, you are very welcome!
Regards,
Joachim