Making a Network from Data with R (Example)
We are back for another tutorial on networks with R. If you missed the introduction or need a refresher, have a look at my previous post to get accustomed to the topic.
Today, we will import a published network that I got the pleasure to work with during my Ph.D. and get it into the graph format using the packages available in R.
In this tutorial, you’ll learn how to import a pairwise list and transform it into a graph network.
The table of content is structured as follows:
Let’s dive into it!
A Small Recap and Some Important Concepts
In my previous tutorial, I mentioned that not all networks are equal. Even for networks that have the same number of nodes and links, they can differ in their directionality. With undirected networks, going from A to B is the same as going from B to A. The adjacency matrix that represents this network in mathematical form will be symmetric.
Networks can also be directed (like the one we will be playing with today). Such a network can represent unilateral relationships (i.e., A to B exists whereas B to A does not) as well as reciprocal relationships (both A to B and B to A are represented).
The adjacency of a directed network will be asymmetric and can thus have different 0s and 1s between relationships A–>B and B–>A. In both cases, nodes can also have relationships with themselves because it does not violate symmetry (i.e., A to A).
Understanding these concepts has certain mathematical implications when it comes to analyzing networks.
For a network with S nodes, it means that:
- There will be S*S possible links for directed networks,
- There will be S*S/2 possible links for undirected networks where nodes are allowed to interact with themselves (some 1s on the diagonal),
- There will be S*S/2 – S possible links for undirected networks where nodes are forbidden to interact with themselves (only 0s on the diagonal). Now that you know a bit more, we are ready to import our first network!
Setup and Get the Data
The data is located at Figshare database. Download the revised pairwise list (the file that weighs 118.44kB) by selecting the file in the list and clicking on the download button.
We will use the classic library igraph for handling networks. This library is both designed to analyze and visualize network graphs. If you are missing the libraries, you can easily install them from the CRAN repository. Check this tutorial if you need a reminder.
library(tidyverse) library(glue) library(igraph)
file_path <- file.path("PairWiseList.txt") # Replace by the path to the file, see ?file.path my_first_graph <- readr::read_delim( file = file_path, delim = "\t", col_types = "cccc", col_names = TRUE)
You might have noticed something: this data does not really look like the adjacency matrix I introduced in my previous tutorial.
The adjacency matrix can also be represented as a paired list, a.k.a. pairwise list (or edge list), that will only contain the links that are present (i.e., the 1s). The pairwise list is more convenient for storage because we only need to store the 1s.
Now that we imported the pairwise list let’s clean up a bit and reconstruct the adjacency matrix. The most important part is to remember that the links have a direction, whether we are dealing with an undirected or a directed network. When using a pairwise list, we need to know which column represents “from” and which represents “to”.
The dataset we are playing with is ‘food web’, a network of food relationships between prey and predators.
It is a directed graph where prey are eaten by predators. Thus the column “PREY” is the “from” and “PREDATOR” is the “to”.
pairwise_list <- my_first_graph |> select(PREY, PREDATOR) # select the columns "PREY" and "PREDATOR"
Let’s now reorder the columns to represent the directionality of links!
We could either use a function from a package or make one ourselves. The function from igraph will automatically handle a pairwise list but often lack flexibility, for instance, if we want to add weights to the links or additional information. The best is to convert our list to a matrix by writing a small helper function.
Before diving in, we need to be aware of a couple of things. First, we need to know the complete list of nodes. Because some relationships can be unilateral, it is possible that some nodes are only represented in the “from” or “to” column. We need to combine the two and remove the duplicates.
prey <- pairwise_list |> pull(PREY) predator <- pairwise_list |> pull(PREDATOR) nodes <- unique(c(prey, predator)) n_nodes <- length(nodes) # 233 nodes
We now know that the adjacency matrix needs to be 233 by 233 and have rows and columns with the names of the nodes. We can build a matrix that will welcome the 1s.
empty_adj_matrix <- matrix( data = 0, nrow = n_nodes, ncol = n_nodes, dimnames = list(nodes, nodes))
Remember that the links are represented by what lies at the intersection of rows and columns in the matrix. We can fill this matrix by scrolling through the rows and columns. Let’s write a function using what we know so far:
pairwise_to_adjacency <- function(from, to) { nodes <- unique(c(from, to)) n_nodes <- length(nodes) if (length(from) != length(to)) {stop("from and to must have same length")} n_links <- length(from) adj_matrix <- matrix( data = 0, nrow = n_nodes, ncol = n_nodes, dimnames = list(nodes, nodes)) for(l in 1:n_links) { a_prey <- from[l] a_predator <- to[l] adj_matrix[a_prey, a_predator] <- 1 } return(adj_matrix) }
adj_matrix <- pairwise_to_adjacency(from = prey, to = predator) sum(adj_matrix) == nrow(pairwise_list) # check if as many links as 1s in the matrix #TRUE
From matrix to graph!
Final step before to enjoy our network: converting the matrix to a graph!
network_foodweb <- igraph::graph_from_adjacency_matrix( adjmatrix = adj_matrix, # see ?graph_from_adjacency_matrix mode = "directed", # directed network weighted = NULL, # no weights diag = TRUE, # 1s at diagonal add.colnames = NULL # colnames of matrix as node attributes )
The package igraph returns objects of class “igraph” which is a special format for storing graphs. The igraph object contains information on which are the nodes (a.k.a. vertices), which are the links (a.k.a. edges), and whatever attributes we want to give nodes and edges (e.g. color, weight, shape).
We can extract information on both nodes and edges using the various functions included in igraph.
vector_nodes <- igraph::V(network_foodweb) # subset the list of vertices/nodes vector_links <- igraph::E(network_foodweb) # subset the list of edges/links
To plot the network, we should simply call plot() on the network object.
plot(network_foodweb)
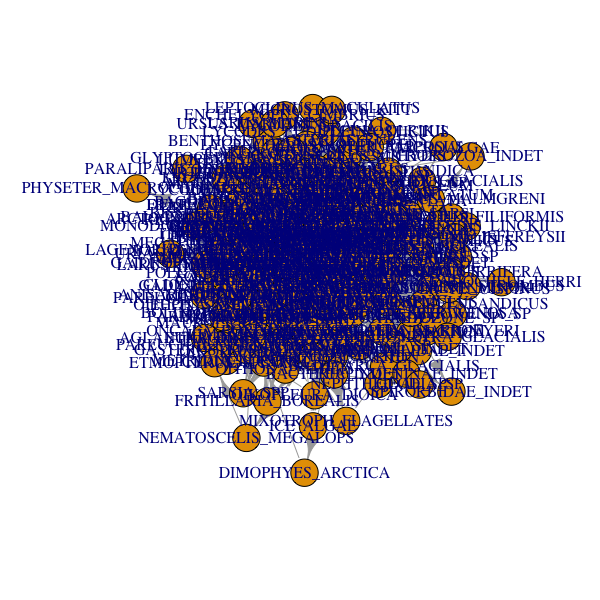
As you can see, the default representation is not ideal. It does not know how to place each node and link and will tend to place nodes that share many links close to one another. They will end up overlapping. Igraph deals with that problem by letting you create layouts.
In the next tutorial, I will show you how to refine the network visualization creating your own layout and custom location for each node and link.
In the meantime, if you want to check some cool igraph tutorials, you can check the blog of Associate Professor Katya Ognyanova.
Video, Further Resources & Summary
Do you need more explanations on how to create a network in R? Then you should have a look at the following YouTube video of the Statistics Globe YouTube channel.
In the video, we explain how to create a network from data in R.
The YouTube video will be added soon.
Furthermore, you could have a look at some of the other tutorials on Statistics Globe:
- Introduction to Network Analysis
- Network Analysis in R
- Network Visualization: Custom Layouts
- Create Symmetric Matrix in R
This post has shown how to make a network from a dataset in R. In case you have further questions, you may leave a comment below.
This page was created in collaboration with Pierre Olivier. You may have a look at Pierre’s author page to read more about his academic background and the other articles he has written for Statistics Globe.
Subscribe to the Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe.
I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Welcome to the Statistics Globe newsletter. From now on, I’ll send you regular emails about statistics, data science, AI, and programming with R and Python.
I’m Joachim Schork. On this website, I provide statistics tutorials as well as code in Python and R programming.
Statistics Globe Newsletter
Get regular updates on the latest tutorials, offers & news at Statistics Globe. I hate spam & you may opt out anytime: Privacy Policy.
Thank you!
Please check your email inbox and click the confirmation link to complete your subscription. If you don’t see the email within a few minutes, please also check your spam/junk folder.







